April 5, 2018 report
Genome sequencing shows baleen whales intermingled more than thought
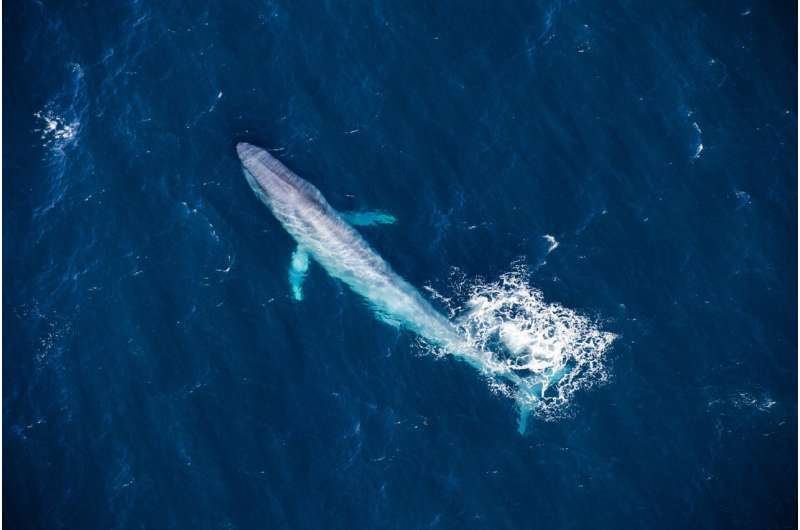

A team of researchers with members from Senckenberg Biodiversity and Climate Research Centre, and Goethe University in Frankfurt am Main, Germany and the University of Lund, in Sweden has found that genetic ties between baleen whales are far more complicated than previously thought. In their paper published on the open access site Science Advances, the group describes their study of the whales using genome sequencing and what they found by doing so.
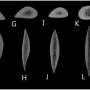
Baleen whales do not have teeth—instead, they have a baleen, consisting of plates of whalebone in the mouth that strain plankton from seawater. Baleen whales include blue, gray, right humpback, rorquals and several others. Because they live in the vast ocean, it is difficult to study whales—scientists have to rely on dead carcasses and the occasional sample taken from a whale when it passes near a research vessel. This has resulted in limited knowledge regarding their biology and evolutionary history. In this new effort, the researchers focused on the history of baleen whales. They obtained six samples from baleens that had been darted and from dead animals, and subjected them to genomic sequencing to learn more about their past.
The researchers soon realized that the history of baleens was far more complicated than thought—so complicated that they were averse to describing their evolution as depicting a family tree, instead preferring to call it a network. This is, they report, because of extensive interbreeding. This was surprising, they note, because of the huge differences in size between different baleens. Blue whales, the largest animal ever to exist on Earth, for example, mixed with the sei, the third largest of all whales, but which is nonetheless dwarfed by the enormous blue. The team reports that they were also able to see that the North Atlantic right whales and bowhead whales had become separate species approximately 28 million years ago.
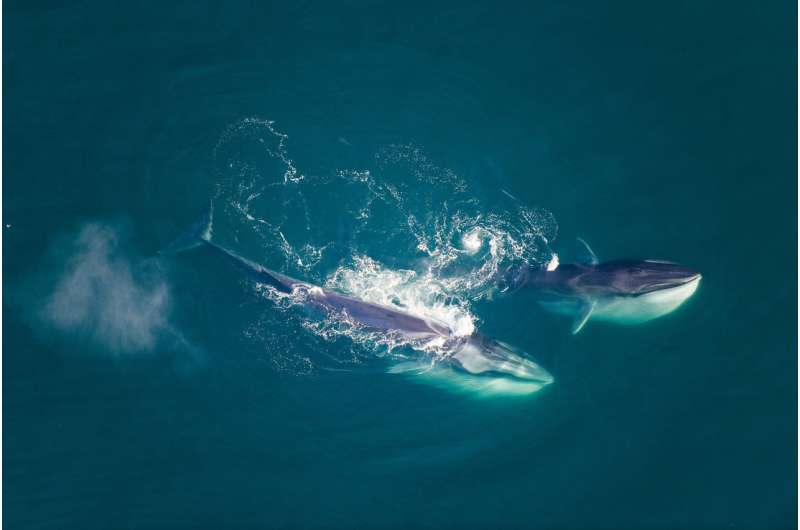

The researchers note that many of the species appeared to evolve to their current state due more to personal tastes, rather than location or geography, which is more often the case with land animals. A taste for tiny creatures obviously drove the evolution of the baleen in general, they add, but differences in which prey they fancied appeared to be the driving factor in the development of shape and size differences.
More information: Whole-genome sequencing of the blue whale and other rorquals finds signatures for introgressive gene flow, Science Advances 04 Apr 2018: Vol. 4, no. 4, eaap9873 , DOI: 10.1126/sciadv.aap9873
Abstract
Reconstructing the evolution of baleen whales (Mysticeti) has been problematic because morphological and genetic analyses have produced different scenarios. This might be caused by genomic admixture that may have taken place among some rorquals. We present the genomes of six whales, including the blue whale (Balaenoptera musculus), to reconstruct a species tree of baleen whales and to identify phylogenetic conflicts. Evolutionary multilocus analyses of 34,192 genome fragments reveal a fast radiation of rorquals at 10.5 to 7.5 million years ago coinciding with oceanic circulation shifts. The evolutionarily enigmatic gray whale (Eschrichtius robustus) is placed among rorquals, and the blue whale genome shows a high degree of heterozygosity. The nearly equal frequency of conflicting gene trees suggests that speciation of rorqual evolution occurred under gene flow, which is best depicted by evolutionary networks. Especially in marine environments, sympatric speciation might be common; our results raise questions about how genetic divergence can be established.
Journal information: Science Advances
© 2018 Phys.org