Duel of the inflammatory master regulators—insights for drug discovery
Anti-inflammatory drugs such as dexamethasone can have harmful side effects on the skin, bones and metabolism. Structural biology research from Emory University School of Medicine has implications for the long-standing quest to separate these drugs' benefits from their side effects.
The findings were recently published in Nature Communications.
Dexamethasone is a synthetic glucocorticoid hormone, used to treat conditions such as allergies, asthma, autoimmune diseases and cancer. It mimics the action of the natural hormone cortisol. Both cortisol and synthetic hormones act by binding the glucocorticoid receptor (GR) protein.
GR can bind DNA in two modes. At some sites, it pairs up or "dimerizes" – turning genes on. At others, it binds one at a time, turning genes off. For GR-targeting drugs, the side effects are thought to come from turning on genes involved in processes such as metabolism and bone growth, while the desired anti-inflammatory effects result mainly from turning inflammatory and immune system genes off.
In their new paper, Eric Ortlund, Ph.D., and colleagues report that GR's ability to directly bind DNA extends more broadly than previously appreciated. The first author is Will Hudson, Ph.D., previously a graduate student with Ortlund and now a postdoctoral fellow in Rafi Ahmed's lab at Emory Vaccine Center.
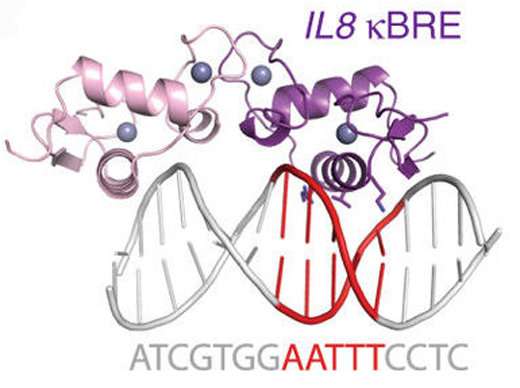
GR was known to interfere with another important DNA-binding protein called NFkB, a master regulator of inflammation. Ortlund's team found that GR can directly bind one at a time to many of the same stretches of DNA that NFkB interacts with.
"This type of interaction, where GR is acting one at a time – we think it's druggable," says Ortlund, who is associate professor of biochemistry at Emory University School of Medicine.
He adds that the paper's findings could lead to the reinterpretation of several studies in the field of inflammatory gene regulation. GR was previously proposed to interact with NFkB sites by "tethering," based on protein-protein interactions.
Ortlund notes that mutations that interfere with the ability of GR to dimerize do not affect its ability to turn down inflammation. On the other hand, mutations that disrupt its ability to bind DNA foil both its activating and repressing functions.
The researchers measured the affinity between GR protein and DNA at NFkB-binding sites and showed that it was similar to other hormone-driven interactions GR was well-known for. They also probed the mode of interaction between GR protein and NFkB -binding sites, using both X-ray crystallography and NMR (nuclear magnetic resonance). They showed that GR binds those sites one at a time, in a region that is actually in between the two stretches of DNA contacted by NFkB itself.
Are the same kinds of interactions happening in cells? Hudson, Ortlund and colleagues re-analyzed data from others to show that direct one-at-a time DNA binding by GR could be responsible for repression of many inflammation genes.
More information: William H. Hudson et al. Cryptic glucocorticoid receptor-binding sites pervade genomic NF-κB response elements, Nature Communications (2018). DOI: 10.1038/s41467-018-03780-1
Journal information: Nature Communications
Provided by Emory University