Proteins date back to the time of sabre toothed cats

Plant scientists at The University of Western Australia have discovered a completely new family of small proteins called PLPs (PawL Peptides) that form by piggybacking on other proteins.
The scientists found 'protein-piggybacking' of PLPs began in seeds of a daisy ancestor that existed 45 million years ago during the Eocene Epoch period—around the time sabre toothed cats walked the earth.
The study included researchers from The University of Western Australia, La Trobe University in Melbourne and the University of California San Diego in the USA and has been published in the open access journal Plant Direct.
The team has previously shown that proteins like these evolved inside an unrelated protein host, but just how big this family was took them by surprise.
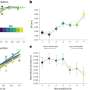
Using instruments capable of separating proteins before weighing them with extreme precision, the team separated and sequenced nearly 50 tiny PLPs. The structures for some were analysed using machines with strong magnetic fields used to align the atoms. Most PLPs consisted of only seven or eight amino acid blocks.
The work will help scientists understand the behaviour of proteins and could be applied in therapeutic drug treatment or even for use in nanotube devices.
Lead author Mark Fisher, Ph.D. student in UWA's School of Molecular Sciences said the research confirmed that PLPs have been around since ancient times and evolved into a huge and diverse class found among members of the daisy family.
"Super-stable peptides like these PLPs are thought to represent the next generation of drugs and we have just found dozens of them and they're unlike anything seen before," Mr Fisher said.
"For a long time, proteins this tiny were assumed to be assembled like LEGO, one amino acid block at a time. We've shown they are chopped out of much larger proteins encoded by genes.
"Their abundance and the way so many sequences are buried inside a host protein has us wondering if other valuable molecules are similarly hiding in proteins from other organisms, are just waiting to be discovered."
UWA Laboratory Head Dr. Joshua Mylne said that the work had unearthed a protein family larger than many others like it.
"For years we had been unwittingly throwing these tiny proteins in the rubbish along with all the fats and oils as we purified the proteins out of seed extracts. They are so small, they hardly behave like proteins at all.
"Now we have so many of them in our hands, it's time to work out why plants are making so many different ones in their seeds. These proteins are only found in the seeds of daisies and it's hard to know what they're doing as they vary so greatly in their properties."
Dr. Mylne led the team of Australian and US scientists that revealed this new family of proteins through the Australian Research Council-supported study "A family of small, cyclic peptides buried in preproalbumin since the Eocene epoch."
More information: Mark F. Fisher et al. A family of small, cyclic peptides buried in preproalbumin since the Eocene epoch, Plant Direct (2018). DOI: 10.1002/pld3.42
Provided by University of Western Australia