Bioinformaticians studied the evolution of broken genes in a fruit fly
Pseudogenes lose their ability to encode proteins, which is unaffected by natural selection, as no mutations in such genes are harmful. A study conducted with the participation of the specialists from the Faculty of Bioengineering and Bioinformatics, MSU (Lomonosov Moscow State University) confirms this, but shows that natural selection still has a certain influence on some of such genes. An article about it was published in the journal Genome Biology and Evolution.
Sometimes, a protein stops playing a key role in an organism's adaptation and survival (e.g. due to environmental changes). In this case, any mutations in the gene that codes this protein become irrelevant for the organism in general.
"Scientists have known for a long time that after a gene undergoes a harmful mutation, e.g., a nonsense mutation, it stops working, and turns into a pseudogene. In this case, negative selection (which removes harmful mutations from the population) stops applying to it, and nonsynonymous mutations (which cause the replacement of one amino acid in a protein with another) start to accumulate in the gene at the same speed as synonymous ones (in which amino acids are not replaced). Previously, this was indirectly referred to in many works with individual examples, but no full-scale research of the effect of nonsense mutations has been ever conducted before," says Nadezhda Potapova, a co-author of the work, postgraduate of the Faculty of Bioengineering and Bioinformatics, MSU.
The human genome contains 53 to 100 non-functioning alleles, and 21 to 27 of them are switched off due to nonsense mutations. This type of mutation got its name due to its ability to deprive the gene of its sense (a functioning protein). Nonsense mutations are a type of mutation causing the occurrence of a terminating codon at which protein synthesis stops. Such mutations have multiple causes: replacements, insertions, or even deletions of the "letters" in the genetic code.
The large number of alleles with nonsense mutations intrigued the scientists. Several molecular mechanisms that allow the function of an apparently broken gene to remain at least partially preserved have already been described. Nonsense alleles differ from nonsynonymous mutations by their harmfulness. They cause diseases three times more often, are observed less frequently, and often have additional copies in the genome (paralogs), making the elimination of just one of them irrelevant.
Biologists studied the genomes of Zambian populations of Drosophila melanogaster that contain 35 nonsense mutations on average. They focused only on the nonsense alleles that occurred due to the replacement of one nucleotide.
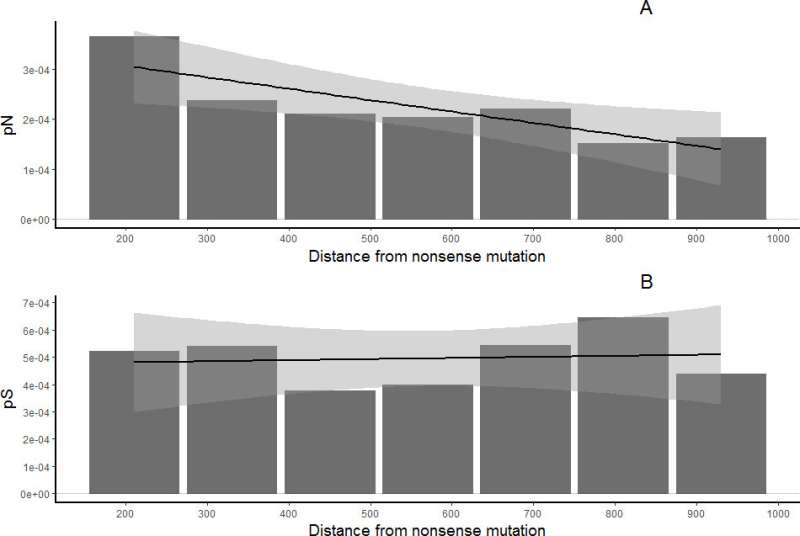
It turned out that negative selection (i.e., the selection reducing the frequency of a certain feature within a population) did not influence those parts of the flies' coding genes) that contained nonsense mutations. However, the selection still had slight impact on some of them. The reason could be alternative splicing—different ways of reading the same DNA region due to the shifting of their start and end points. Biologists believe that alternative splicing is the reason that some genes preserve their functions and at least partially exposing them to negative selection.
"We used Drosophila melanogaster genomes. We managed to confirm that nonsense mutations actually switch off single exon genes (i.e. the genes that contain no introns cut out during protein synthesis), and they start to accumulate both synonymous and nonsynonymous mutations at the same speed. In multi-exon genes (genes with several introns) only the exon with the mutation seems to be left out, and the others remain under the influence of negative selection, though slightly weakened," explains Nadezhda Potapova.
More information: Nadezhda A Potapova et al, Are Nonsense Alleles of Drosophila melanogaster Genes under Any Selection?, Genome Biology and Evolution (2018). DOI: 10.1093/gbe/evy032
Provided by Lomonosov Moscow State University