Team uses magnetic beads to model microscopic proteins, polymers

An engineered string of micronwide beads may take up the slack where computer modeling fails researchers who study the bending, folding and other movements of polymers or biomolecules like actin and DNA.
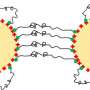
Rice University chemical and biomolecular engineer Sibani Lisa Biswal and her students—lead author Steve Kuei, a graduate student, and co-author Burke Garza, an undergraduate—created strings of polystyrene beads enhanced with iron to magnetize them and with streptavidin, a natural protein that serves as a springy linker between them.
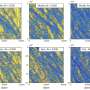
They placed the strings into solutions and manipulated them with a rotating magnetic field. Some strands were made to be stiff, some a little bendy and some much more flexible. By applying an outside magnetic force, the researchers were able to see how each type of string reacted and compared the results with computer models of strings that had the same properties.
Biswal said the new platform lets researchers study how strings of various types behave under dynamic conditions in a scalable way that isn't possible with simulations because of the high computational cost. It could benefit researchers who study proteins, DNA and RNA in biological systems or those who study the fluid properties of polymers that entangle to create gels or the ordering and packing density of liquid crystals.
"I can see people using this to study the practicalities of building, say, micro robots with wagging tails, or robots that can coil up," Biswal said. Because the technique could model flagellar motion in a fluid environment, it might also help make artificial organisms possible, she said.
The research appears in the American Physical Society journal Physical Review Fluids.
The Rice team knew there was already plenty of information available about rigid and flexible strings, filaments and fibers and how they moved due to Brownian motion or in response to shear or other forces. But there was very little data about semiflexible fibers like actin, carbon nanotubes and cilia.
"There's a lot of interest in materials that fold into complex geometries, but even simple things like tying a knot at the macroscale are very difficult at the microscale," Biswal said. "So we developed a method to let us look at the dynamic forces involved. The ability to engineer different flexibilities into this material is its real power."
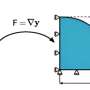
The strings isolated in liquid could be shaken or stirred, but the Rice team built a device to rotate the magnetic field that touched each bead with gentle force. They observed strings that reacted in different ways depending on the level of built-in flexibility and/or elasticity.
Rigid rods simply rotated in concert with the magnetic field. Those with a bit more flexibility "wagged" their tails in the moving field, and the centers would turn as the tails relaxed. Longer and more flexible strings were prone to coiling, eventually compacting into a form with less drag that allowed them to behave like their rigid brethren.
"Most of the time strings are open in structure until you turn on the rotating field and they crumple up," Biswal said. "That changes the underlying fluid properties, because they go from taking up a lot of space to taking up very little. A fluid with strings could go from behaving like honey to behaving like water."
Such effects can't be seen directly with proteins that are both several orders of magnitude smaller and yet have too many beads - the residues - to simulate their folding easily, Biswal said.
"There's been some work with fluorescently labeled DNA and other biofilms like actin, but they can't get that bead-to-bead resolution that we can with our method," she said. "We can actually see the positions of all our particles."
Strings in the current study had up to 70 beads. The researchers plan to make chains up to 1,000 beads for future studies on more complicated folding dynamics.
More information: Steve Kuei et al, From strings to coils: Rotational dynamics of DNA-linked colloidal chains, Physical Review Fluids (2017). DOI: 10.1103/PhysRevFluids.2.104102
Provided by Rice University