Apple genome sequence helpful to breeding of new varieties published

A high quality genome sequence of apple is published in this week's Nature Genetics by an international team of scientists, among which researchers of Wageningen University & Research in the Netherlands. The publication of the sequence facilitates faster and more targeted breeding of new apple varieties with increased disease resistance, improved production traits, and better fruit quality. With this the results support a more sustainable production of apple fruit, both from an environmental and a financial perspective.
The genome sequence was assembled by an international consortium of research institutions from France, Italy, Germany, the Netherlands and South Africa. The high quality of the genome data, indicating over 42 thousand putative genes, is the result of the use of latest sequencing technologies, which generate long stretches of DNA sequences, a very specific apple variety, and the most informative genetic linkage map in apple developed in earlier research.
The genome sequence gives new insights into the organization of the apple genome. Already 93 percent of the 42,000 putative genes have been validated through RNA sequencing. This knowledge is useful for the identification of genes that control a trait of interest and for the development of DNA-based diagnostic tests that can accelerate breeding of new varieties.
The use of a so-called di-haploid apple variety was critical for the success of this study. Apple is an outcrossing species, making its genome heterozygous. Also, apple originated from a hybridization between two different species, which was coupled with a whole genome duplication. As a result, each regular apple variety has up to four variants for each of its DNA sequences. The di-haploid variety used in this study is special as it has only up to two variants of every sequence. This leads to a dramatic complexity reduction, which made it possible to generate a very high quality genome sequence.
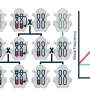
The new insights in the apple genome include a clear view on the duplication patterns among the 17 chromosomes of apple. This facilitates the identification of gene copies with similar function. Next, so called 'repetitive regions' have been assembled. These thus far uncharacterized regions of the apple genome may be involved in regulating gene expression. Finally, a new type of repeat sequence was found that may be specific for centromeres, which may lead to new insights in chromosome division and replication.
The research was coordinated by Etienne Bucher of INRA-Angers. Researchers of Wageningen University & Research contributed to the genome sequencing, genome mapping and assembly, applying their experience and skills in bioinformatics and by giving early access to a high quality reference genetic linkage map in apple.
Wageningen University & Research itself develops new apple varieties, which resulted in the top-variety 'Elstar', 'Santana' and the recently released 'Natyra'. The latter two are suited for biological production since these varieties have disease resistances. Additionally, 'Santana' is suitable for consumption by most individuals with a mild apple allergy.
Wageningen will use the new insights in the DNA of the apple in the targeted breeding of new varieties.
More information: High-quality de novo assembly of the apple genome and methylome dynamics of early fruit development, Nature Genetics (2017). nature.com/articles/doi:10.1038/ng.3886
Journal information: Nature Genetics
Provided by Wageningen University