Breakthrough in 'amphibian plague': Deadly fungus genes identified
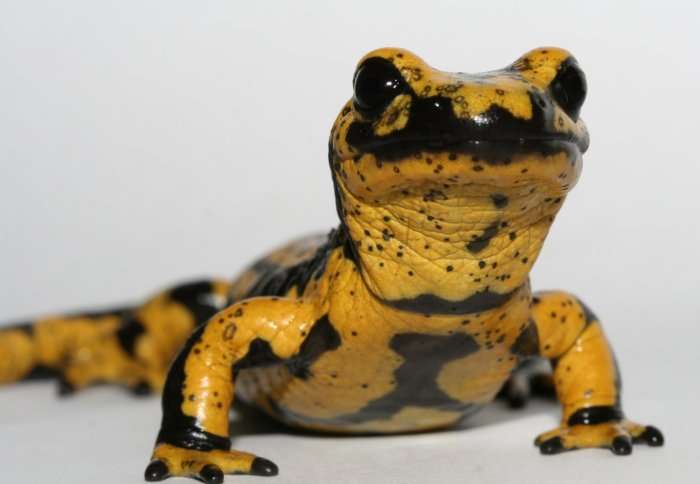

Scientists have identified the genes of a deadly fungus that is decimating salamander and newt populations in Northern Europe.
Batrachochytrium salamandrivorans (Bsal), dubbed the 'amphibian plague', is a highly infectious chytrid fungus that affects many species of salamanders and newts, literally digesting their skin, which quickly leads to death. Since its discovery in 2013, very little has been found about how the fungus causes disease.
Now, researchers from Imperial College London, Ghent University, and the Broad Institute, have sequenced and identified the genes responsible for Bsal from an infected salamander. The authors say the findings, published last week in the journal Nature Communications, could ultimately help conservation efforts and provide drug targets in the future to help curb the disease.
Dr Rhys Farrer, co-author from Imperial's School of Public Health, said: "Until now, no one knew the exact mechanisms Bsal uses to cause disease. Our findings mean that policy makers and conservationists are now equipped with more knowledge on how best to curb this amphibian plague."
Dr Rhys Farrer and co-author Professor An Martel from Ghent University sequenced the genes from a salamander that had died from Bsal, and compared the genes with those of Batrachochytrium dendrobatidis (Bd), a closely related deadly fungus that affects not just salamanders and newts, but all amphibians. Bd has caused more extinction events than any other infectious disease known to science.

The researchers in this study found that, like Bd, Bsal is equipped with a range of genes for infecting and causing disease in amphibians. They found most genes were found in both species, but some genes were uniquely found in either Bd or Bsal. Therefore, the researchers hope that by comparing the genes in common between the two fungus types, they can gain insight of how Bsal works.
One gene category that Bsal and Bd had in common were genes for metalloproteases, which are enzymes that digest amphibian skin so the patches can be colonised by the fungus. However, they were present in vastly different quantities in each type of fungus. This might explain the differences and similarities in the way each fungus type infects amphibians.
For example, Bsal had three times the amount of genes for metalloproteases, which are enzymes that digest amphibian skin so the resulting patches can be colonised by the fungus. This likely explains why skin ulceration is a major symptom of Bsal, but not so much in Bd.
The researchers also found that the newly identified fungal genes are highly 'switched on' during infection of salamanders, but recognise that more work is needed to definitely categorise them as harmful or non-harmful. Unlike Bd, Bsal also appeared to suppress salamanders' immune systems, suggesting that this is a mechanism through which Bsal colonises its hosts.

Dr Farrer said: "The genes we have sequenced are from only one infected salamander, and as such, there could be variation in the population of salamanders and Bsal that affects their interactions. Furthermore, no drug has currently been identified to target the genes we found, or the proteins they help create. We hope to eventually know enough about how Bsal infects their hosts to be able to target and treat those areas."
Bsal was first discovered in 2013 by scientists from Imperial College London and Ghent University who were investigating a massive drop in numbers of fire salamanders in the Netherlands. The disease appears to have originated from Asia and is now spreading through Europe. Last year, the US government banned the import of salamanders to reduce the threat posed by Bsal, where it hasn't yet been detected.
Dr Farrer added: "We are pleased to have finally found some clues about this devastating and rapidly emerging plague."
The next step for the researchers will be to sequence genes from more infected salamanders to build a bigger picture of the genes.
More information: Rhys A. Farrer et al. Genomic innovations linked to infection strategies across emerging pathogenic chytrid fungi, Nature Communications (2017). DOI: 10.1038/ncomms14742
Journal information: Nature Communications
Provided by Imperial College London