'Neighbor maps' reveal the genome's 3-D shape
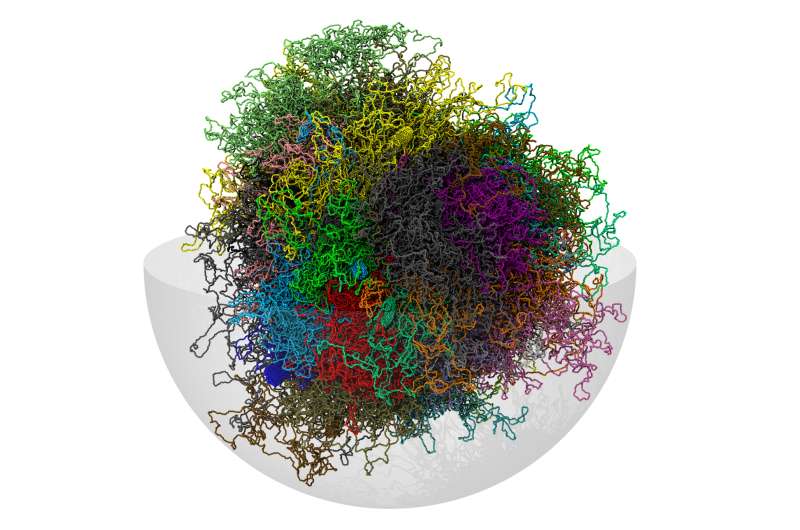
A group coordinated by SISSA Trieste has built a 3-D computer model of the human genome. The shape of DNA (and its sequence) affects biological processes and is crucial for understanding its function. The study has provided an approximate but realistic 3-D identikit of the human genome. The resulting structural reconstruction, based on experimental information and statistical methods, will be refined as new experimental data become available. The study is published in Scientific Reports.
Genome sequencing is a milestone in modern biology that allows access to the entire genetic sequence for the development and function of organisms. Sequencing the genome is a bit like writing down the exact order of the colour of beads in a necklace—knowing how they are arranged along the thread gives no indication regarding the shape of the necklace. The shape of the DNA strand can be highly complex, given that the chromosomes are loosely arranged in an apparently chaotic tangle in the cell nucleus. Since the shape of chromosomes may have a decisive effect on their function, it is important that it should be characterised, in part because scientists think the DNA tangle in the nucleus is only apparently chaotic and that it has instead a specific "geography" for each tissue and stage of cell life.
"Arriving at a precise description of the shape of the DNA tangle is unfortunately incredibly complicated," explains Cristian Micheletti, SISSA professor and coordinator of the new study. "In our case, we used experimental data on 'proximity pairs.'"
"Imagine having to create a map of a city based only on information like 'the post office is opposite the station,' 'the chemist is close to the gym,' 'the fruit and vegetable market is near the football field,' and so on. If you have only a small number of such statements to go by, your map will be approximate, and in some cases indeterminate. But if you have hundreds, thousands or even more, then your map will become increasingly precise and accurate. This is the logic we followed."
"Proximity pairs" therefore refer to information on the closeness of two points on the map. In the case of nuclear DNA, this information was provided by a technique known as Hi-C, developed by North American research groups in 2010. In this chemical-physical technique, bits of genome located close to each other in the nucleus are tied together and identified by sequence. By collecting large numbers of these proximity pairs, scientists discovered which points of the chromosomes lie close to each other in the nucleus. While this is today the most powerful technique for investigating DNA organisation in the nucleus, it is still inadequate for inferring its overall shape. "For this reason, we thought we would try to go further," says Micheletti.
More information: Hi-C-constrained physical models of human chromosomes recover functionally-related properties of genome organization. Scientific Reports DOI: 10.1038/srep35985
Journal information: Scientific Reports
Provided by International School of Advanced Studies (SISSA)




















