New design of large-scale microparticle arrays for bioengineering applications

Arrays of microparticles are common in many material science and bioengineering applications, but can be tedious to use because of their limited capacity. Large-scale microparticle arrays (LSMAs) can make analysis more efficient and precise, allowing for the placement and study of many items at once. Unfortunately, today's techniques of moving to a large-scale platform cannot simultaneously accomplish the requirements of scalability, precision, specificity and versatility that would make the use of LSMAs practical.
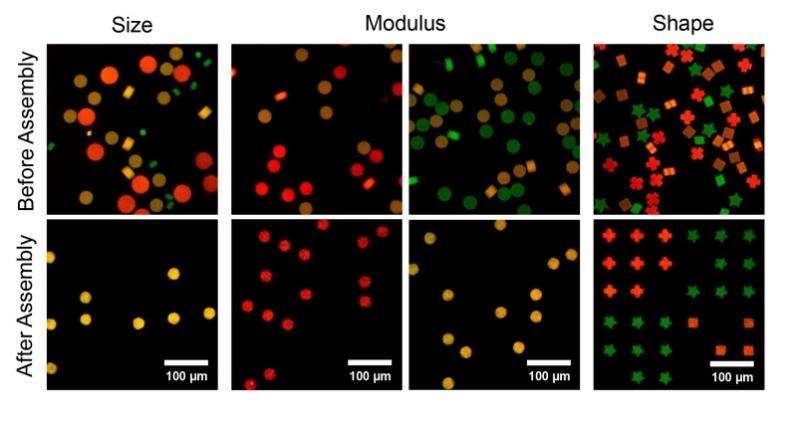
Researchers from MIT and the Massachusetts General Hospital (MGH) have developed a new technique using porous microwells that pushes the precision and scalability of LSMAs to a new extreme. This new method, described in the Sept. 5 issue of Nature Materials, uses fluid flow to guide tens of thousands of microparticles at once, pushing them into microwells as the fluid moves through small open pores at the bottom of the porous well arrays. The new LSMA technique sorts and arrays particles on the basis of their size, shape, or modulus. This sequential particle assembly allows for contiguous and nested particle arrangements, as well as particle recollection and pattern transfer.
"Today's applications are increasingly complex; this new technique creates the most precise arrangement of particles, allowing for a more detailed and accurate array," says Patrick Doyle, the Robert T. Haslam (1911) Professor of Chemical Engineering and Singapore Research Professor at MIT. "This technique opens the way to new applications, including the study of diseased cells and anti-counterfeiting practices."
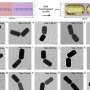
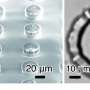
Led by Doyle and Daniel Irimia, associate director of the BioMEMS Resource Center at MGH, the team developed this porous microwell platform where guided microparticles are inserted into congruent microwells, whereas geometrically mismatched particles are removed in a washing step. "Microwells have been used as an assembly template in the past, but they were useful only for single-particle arrangement," says Irimia. "Scaling-up efforts resulted in particle arrangements with some degree of randomness. In our technique, controllable driving forces allow for the positioning of tens of thousands of particles with high specificity."
Bioengineering applications
The ability to generate large arrays of cells is important for cell-screening applications, which aid immunology and the fight against cancer. For example, arranging cells in 2-D arrays for the study of cellular processes that progress over time has significant advantages compared with serial approaches, such as flow cytometry. Cells in a 2-D array can be analyzed more than once and several cells imaged simultaneously. This creates a higher yield which helps to avoid potential differences between the first and last cell analyzed.
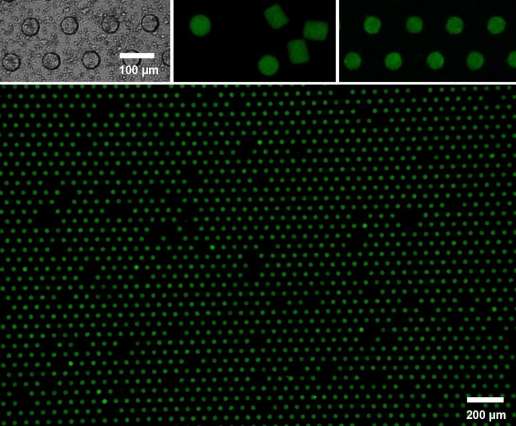
The team tested the performance of LSMA techniques by generating arrays of more than 10,000 mammalian glioma cells, which can cause brain tumors. An acceptable yield for each array took approximately 60 seconds, significantly faster than the previous method using passive cell settling in microwells, which requires between five and 40 minutes. "Because of the speed in which the cells are arranged, there is little change in the cell's state," says Doyle. "This gives us more time to observe how cells respond to drugs and disease."
This research began when a team at MGH, led by Irimia, attempted a new way to arrange cells for analysis. "Understanding how neutrophils [the most abundant type of white blood cells in mammals] react to stimuli helps us to understand how inflammation starts and evolves inside the body," he says, "This technique allows a level of complexity we've not had before; we can analyze the same cells repeatedly and therefore gather more information about their function and interactions in less time."
Anti-counterfeiting applications
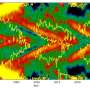
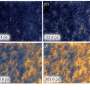
The team also demonstrated that this new technique is compatible with particle recollection and pattern transfer. To demonstrate the encoding/decoding capacity of LSMAs, the researchers generated a 2-D arrangement of nanocrystal-laden microparticles for use in anti-counterfeiting. These nanocrystals, developed by Doyle and his team specifically for use in anti-counterfeiting applications, glow when exposed to near-infrared light. They can be altered to emit any color, allowing for the creation of unique barcodes invisible to the naked eye.
Conventional printing of the microparticle barcodes resulted in limited precision and resolution. By using a prealigned microwell array, this new approach generates a high-resolution, multicomponent pattern. The pattern is then transferred to a target object, like a poker chip. In tests, an image of the transferred pattern was taken with an iPhone under near-infrared exposure and was successfully decoded within 10 seconds.
"This development can impact future precision medicine since the platform can be effectively applied in many precision high-throughput molecular diagnostics, single cell analysis, and other innovative quantitative cell biological experiments," says Luke Lee, the Arnold and Barbara Silverman Distinguished Professor of Bioengineering, Electrical Engineering and Computer Science, and Biophysics at the University of California at Berkeley, who was not involved in the research. "Since smart microparticle technology with barcodes has great potential in life sciences and clinical applications, this team's new solution for scalability is a great accomplishment for a large-scale automated precision biology and medicine."
More information: Jae Jung Kim et al. Porous microwells for geometry-selective, large-scale microparticle arrays, Nature Materials (2016). DOI: 10.1038/nmat4747
Journal information: Nature Materials
Provided by Massachusetts Institute of Technology
This story is republished courtesy of MIT News (web.mit.edu/newsoffice/), a popular site that covers news about MIT research, innovation and teaching.