Structure of key DNA replication protein solved

A research team led by scientists at the Icahn School of Medicine at Mount Sinai (ISMMS) has solved the three-dimensional structure of a key protein that helps damaged cellular DNA repair itself. Investigators say that knowing the chemical structure of the protein will likely help drug designers build novel anti-cancer agents.
The study, published in the October 21 issue of the journal Science Advances, involved a team of investigators from multiple institutions, who worked for more than two years to decipher the unusual configuration of the protein PrimPol, whose function was discovered in 2013. PrimPol is used in cells when normal repair proteins encounter damaged sections of DNA, often caused by anticancer chemotherapy drugs. The protein can skip over the damage to rescue DNA replication, says the study's senior investigator, Aneel K. Aggarwal, PhD, Professor of Pharmacological and Oncological Sciences at ISMMS.
"PrimPol can counter the anti-cancer action of common chemotherapeutic agents such as cisplatin. By inhibiting PrimPol, we believe that we can increase the efficacy of chemotherapeutic agents in the treatment of many cancers," he says.
DNA damage happens constantly—more than 100,000 events occur in every human cell each day. PrimPol is necessary for the cell to repair DNA damage, but sometimes this may not be to the individual's benefit, as in the case of resistance to chemotherapeutic agents, says the study's co-lead author, Olga Rechkoblit, PhD, Assistant Professor of Pharmacological Sciences at ISMMS.
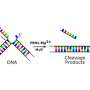
The basic steps involved in DNA replication are known. The first step involves unzipping the intertwined double helix DNA structure, creating a "Y" shape known as a replication fork. These two strands act as templates for making the new DNA strands. A short piece of RNA known as a primer (produced by a primase enzyme) acts as the starting point for the synthesis of new DNA.
"It had been believed that DNA polymerase and primase activities in human cells were the province of separate enzymes. Then PrimPol was discovered, and the understanding of DNA replication changed dramatically. PrimPol was found to be capable of both restarting and performing DNA synthesis after DNA replication stalls," says Dr. Rechkoblit.
While scientists were excited by the discovery of the enzyme, they didn't understand how it worked. Drs. Aggarwal and Rechkoblit organized a team of investigators from Cornell University in New York, Argonne National Laboratory in Illinois, and the University of Texas Medical Branch in Galveston to find out.
"Because the three-dimensional structure of the enzyme is so different from that of other DNA polymerases, it required a group effort to elucidate its structure," says Dr. Aggarwal.
The clinical implications are clear, Dr. Rechkoblit says.
"Many chemotherapy agents kill cancer cells by damaging their DNA and preventing the completion of DNA replication. PrimPol, on the other hand, promotes the replication progression and, thus, cell survival," she says. "Knowing the structure of PrimPol described in the current study is invaluable for designing an inhibitor for this enzyme as a future cancer therapy."
"The idea would be that a patient could be given a PrimPol inhibitor—an agent that shuts down this clever machine—at the same time as chemotherapy, so that cancer cells cannot repair the killing damage chemotherapy offers," she says.
More information: O. Rechkoblit et al, Structure and mechanism of human PrimPol, a DNA polymerase with primase activity, Science Advances (2016). DOI: 10.1126/sciadv.1601317
Journal information: Science Advances
Provided by The Mount Sinai Hospital