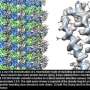
Cryo-EM's renaissance: NERSC resources help illuminate human molecular machinery
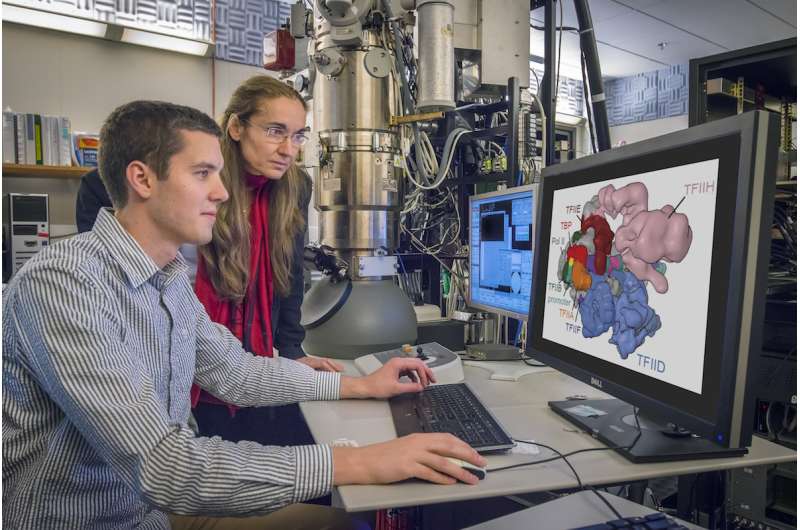

In a pair of breakthrough Nature papers published recently, researchers in Eva Nogales' Lab at UC Berkeley and the Lawrence Berkeley National Laboratory (Berkeley Lab) mapped two important protein functions in unprecedented detail: The role of TFIID, effectively improving our understanding of how our molecular machinery identifies the right DNA to copy; and how proteins unzip double-stranded DNA, which gives us insights in to the first-key steps in gene activation.
They captured these processes at near-atomic level resolution by freezing purified samples and photographing them with electrons instead of light, a process called cryo-electron microscopy (cryo-EM). The researchers then used supercomputers at the National Energy Research Scientific Computing Center (NERSC), a Department of Energy (DOE) user facility, to process and analyze the data.
They note that these papers are representative of the renaissance currently under way in the cryo-EM field—driven primarily by the rise of cutting-edge electron detector cameras, sophisticated image processing software and access to supercomputing resources and expertise at facilities like NERSC.
"This is a revolutionary time for cryo-EM," says Eva Nogales, a researcher at both UC Berkeley and Berkeley Lab. "One advantage of this technique is that it doesn't require researchers to crystallize samples, which is a huge bottleneck. Combine this benefit with the rise of new detector technology that can image molecular-sized structure at very high-resolution and now you have everyone's attention."
The new technology has inspired researchers who traditionally have not been interested in it—like crystallographers and biochemists—to work with cryo-EM, she adds. Biologists are now motivated to investigate even more questions, many of which were previously impossible to answer.
"It's just incredible," says Nogales. "Today there are more investigators in the field, and the number of projects that each of us wants to tackle is growing exponentially. So now there is more data per project, more projects per lab and a lot more labs doing this work. This is leading to an explosion of data."
The data explosion is also compounded by the fact that new electron detector cameras can now take movies—essentially stacks of images. Yuan He, a postdoctoral fellow who previously worked in Nogales' Lab and is now an assistant professor at Northwestern University, notes that this capability is what allows high-resolution information being extracted from practically every image taken with the new cameras.
"When electron microscopes put high-energy electrons through your sample, that sample is going to move. If you are just taking single images, the movement can cause a blurring effect in your data that is very hard to correct because the motions are inevitable and random," says He. "But if you are collecting data in a movie format, you can correct the blurring and get a high-resolution image by realigning the movie frames to each other."
"When we were collecting data as single images, each data set was on the order of 100 gigabytes. Now that we are collecting movies, our datasets have increased up to 100 fold," says Robert Louder, a biophysics graduate student in Nogales' Lab.
Supercomputing Shapes the Future of Cryo-EM
As the data sets grow in size, so does the demand for supercomputing resources to process this information. "The data processing for cryo-EM is not trivial," says Nogales. "Our analysis requires a lot of human intervention, we frequently adjust parameters on the fly and can't be sitting in long queues."
So she and her team worked with NERSC Engineer Joaquin Correa and the facility's user consultants to come up with a queue strategy that would work for their research. They also worked with other NERSC staff to get their codes to scale efficiently and effectively on the facility's supercomputers.
"The new cameras allow us to look at structures with much higher detail, at much higher resolution, but it takes much more computational power to process the images than what we have used in the past," says Louder.
Cryo-EM researchers used to be able to run their analysis programs on desktop workstations and get results in one or a few days, he adds, but with today's more sophisticated processing programs researchers using desktop computers would be waiting more than a week. This is a problem because processing cryo-EM data is an iterative process, and researchers need quick feedback to test different parameters.
According to Louder, this is where access to NERSC resources came in handy. By running their jobs on NERSC systems the team could execute their large jobs on hundreds of processors, run multiple instances of these jobs and compare results without waiting. They were able to get results on the order of hours or maybe a day.
"Access to computation is still a limitation for many labs, but access to NERSC means that computation wasn't a limitation for our lab," says Louder. "Our analysis programs are very I/O intensive, so they need to access large amounts of data rapidly and many times throughout the runs. That is one optimization that we've been working with NERSC staff to tweak. This collaboration has saved us a lot of time, which we can now spend doing more science."
According to Nogales, researchers in the field are currently limited by access to microscopes. But as academic, government and private institutions invest in instruments, she suspects that the next big limitation will be access to supercomputers for data processing.
"This is an exciting time to be using cryo-EM," says He. "It was Nature Method's 2016 'Method of the Year,' and we are seeing high-impact publications of cryo-EM results practically every week."
More information:
Related:
1. phys.org/news/2016-05-unveil-i … a-transcription.html
2. phys.org/news/2016-03-secrets- … s-major-advance.html
Journal information: Nature
Provided by Lawrence Berkeley National Laboratory