Cloud-based resource to support microbiology data sharing
The third week of July 2016 saw the launch of the UK's Cloud Infrastructure for Microbial Bioinformatics (CLIMB)— probably the largest dedicated microbial bioinformatics resource in the world. CLIMB is a collaboration between academic and computing staff at the Universities of Bath, Birmingham, Cardiff, Swansea and Warwick.
The impressive scale of the CLIMB computational infrastructure matches its ambitions as a national resource for the UK's medical microbiology community and our international partners. However, size isn't everything—CLIMB also represents a user-friendly, one-stop shop for sharing software and data between medical microbiologists in the academic and clinical arenas.
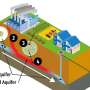
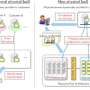
An exciting feature of CLIMB is our use of "cloud computing". This means that rather than dozens, or even hundreds, of research groups across the country having to set up and maintain their own servers, users can access shared pre-configured computational resources on demand. Key to this set-up is the concept of virtualisation, which allows users to work in simulated computer environment populated by virtual machines (VMs), which sit on top of the physical hardware, but look to the user just like conventional servers. Perhaps a useful analogy is with the digitisation of images or text, which allows an object to be stored and reproduced indefinitely without loss of quality.
Virtualisation is widely used to consolidate many VMs on to a single physical machine, ensuring efficient use of hardware. However, this approach has several other important advantages. As with other digital objects, virtual machines can be stored and shared with ease. Thus, you can launch a VM, install programs and pipelines tailored to microbial bioinformatics and then take a snapshot or image of the customised virtual server. These snapshots can then be stored, copied and shared with others in the research community, freeing downstream users of the hassle of installing complex programs and their often-troublesome dependencies.
An example here is CLIMB's use of the Genomics Virtual Laboratory (GVL), which comes preinstalled on one of our standard VM offerings. GVL, which was developed in Australia, provides access via the web and command line to a virtual desktop, a personal Galaxy server, together with fully featured environments for the programming language R and the iPython Notepad. Crucially, GVL also provides access to Torsten Seemann's groundbreaking Nullarbor pipeline, which incorporates a range of microbial bioinformatics analyses, while presenting results as a simplified web report .
However, it is important to stress that the advantages of virtualisation go beyond mere efficiency. The creation of multiple standardised VMs with preconfigured software and settings will facilitate training in microbial bioinformatics. Furthermore, the ability to encapsulate a complex server environment into a publishable digital object will allow the entire community to explore and exploit published pipelines and guarantee that complex bioinformatics analyses can be replicated, enhancing the reproducibility of science.
All this would mean nothing if the system were a private playground for the CLIMB investigators. Instead, CLIMB has been set up as a national facility, with access provided free-of-charge to academic medical microbiologists and microbial bioinformaticians within the UK. But we see CLIMB as more than an academic facility: instead, we hope it will act as a bridge between academics and public health professionals, facilitating sharing of skills, knowledge and approaches between the two communities, as well as exchange of software and data. We are pleased to note that the system has already seen early exploratory use by Public Health Wales, Public Health England and the Animal and Plant Health Agency. We anticipate that our virtualisation approach will facilitate the development of standard operating procedures in clinical and public health microbiology, suitable for national or international accreditation.
Provided by University of Bath