Researchers using blowflies to sample mammal DNA and develop a mammal monitoring tool

Mammal DNA detection in invertebrate guts has been proposed to study mammal diversity but certain concerns remained. The concerns were addressed by showing mammal DNA persists in blowfly guts for 1-4 days post-feeding and developing primers that bind to mammal DNA barcodes that can distinguish lookalike mammal species,but do not bind to blowfly DNA.
According to the Global Mammal Assessment, a program that collects data on the status and threats of mammal species worldwide, most mammal species in the tropics are either threatened or poorly known. The lack of knowledge about tropical mammals could be attributed to the inefficient and labour-intensive tools used for their detection and study. The traditional tools for mammal detection include deciphering footprints and scat encountered while trekking through the forest, and also using wire cages and nets. The latter are becoming less acceptable from an ethics perspective. A recent addition to the mammal monitoring toolbox is camera trapping. But camera traps need clear photos. Unlike humans, some other animals can find other mammals in the forest much more easily especially when those mammals are their prey. With modern advances in DNA sequencing technology, detecting the presence of forest mammals based on DNA detection in the guts of invertebrates has become a potential approach to study the diversity of forest mammals.
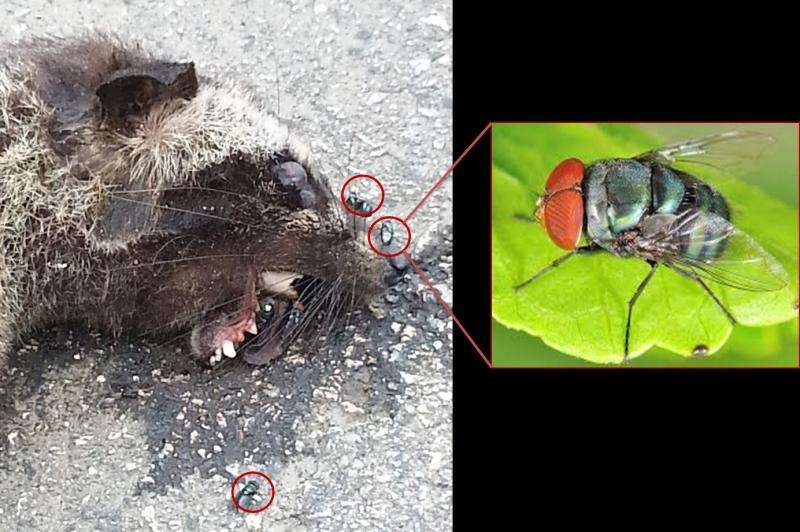
Blowflies - the annoying flies buzzing around cooked meat on dining table as well as faeces, wounds or carcasses of wild animals in tropical forests, may do a better job of sampling mammal DNA than other invertebrates due to their non-selective feeding habits in all types of habitats. Their ability to concentrate on faecal DNA while feeding is another bonus. Chrysomya megacephala, a blowfly known as the oriental latrine fly (pictured), is among the first and most abundant species of invertebrates arriving at mammal carcasses in tropical forests of peninsular Malaysia.
The DNA of the mammals consumed by blowflies is normally fragmented into small pieces during the ingestion process. This raised two major concerns about the suitability of blowfly-derived DNA as a mammal monitoring tool. They are: How long does mammal DNA persist in the guts of blowflies? Which particular fragment of mammal DNA should be targeted to maximize the chances of successful detection?
To address the first concern, a colony of blowflies was reared in the lab at the Museum Zoology, University of Malaya. Blowflies were starved for a fixed period of time, before their guts were dissected. The experiment showed that cow DNA persists in the guts of adult blowflies for one to four days post-feeding. This suggests that DNA should be extracted for blowfly guts within one day of capture in order to detect mammal DNA of sufficient quantity and quality.
To address the second concern - Which DNA fragment to target? The researchers chose to design primers targeting the DNA barcode fragment of mitochondrial DNA that can distinguish lookalike mammal species. The research team designed primers (tags) that bind to a short fragment of the mammal DNA barcodes but didn't bind to the DNA of the blowflies. These primers allow the DNA to be copied (in a process known as PCR), to build up a sufficient number of copies that can be read by a DNA sequencer. The DNA barcode target region amplified by the primers is short enough to be detected among the fragments of ingested mammal DNA and to be sequenced by high-throughput DNA sequencers. However, it contains enough information to be able to distinguish even those lookalike mammal species.
Currently, researchers at the Museum Zoology, UM are comparing the successful mammal detection from guts of blowflies collected in forest with mammals detected in cage traps, mist nets, camera traps, hair traps and scat samples.
More information: Ping-Shin Lee et al. Reading Mammal Diversity from Flies: The Persistence Period of Amplifiable Mammal mtDNA in Blowfly Guts (Chrysomya megacephala) and a New DNA Mini-Barcode Target, PLOS ONE (2015). DOI: 10.1371/journal.pone.0123871
Journal information: PLoS ONE
Provided by University of Malaya