October 29, 2015 report
Researchers find a way to selectively add saryl groups to amino acid residues in proteins
(Phys.org)—A team of researchers at MIT has found a way to modify proteins using palladium reagents, allowing for the possible creation of a host of therapeutic drugs. In their paper published in the journal Nature, the team describes their technique, why it may be useful and applications that may benefit from its use. Heather Maynard with the University of California offers an overview of the work done by the team in a News & Views piece in the same journal issue.
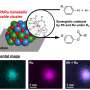
Biological scientists are interested in modifying proteins because doing so can cause them to work differently in the body, by increasing the time a medicine remains working, for example, or by helping to target certain cells, such as those that are part of a tumor. But modifying a protein without disrupting its ability to function is challenging, as Maynard notes—current approaches generally lead to low yields. One approach, using organometallic reagents has been tried, but has not really caught on due to the strict conditions under which reactions must occur. In this new effort, the researchers have found a way to use such reagents without the restrictions, opening the door to it use. With their technique the organometallic reagents react specifically with thiol groups of free cysteines (amino acids) across a broad pH range. More specifically, the reactions they caused used a palladium reagent to marry an aryl group onto a cysteine residue. Such reactions occur very rapidly, they note, disallowing other reactions from occurring—the results have been shown to be very stable as well, allowing for the possible creation of kits that can be sold to chemists and stored for periods of time.
To test their approach, the researchers modified several proteins in different ways, focusing on staple peptides and conjugates—proteins that are useful as therapeutic drugs. They report that they were able to modify one such protein in just ten minutes, with 100 percent yield. The process still needs refining Maynard notes—such as reducing the amount of palladium needed, because the amounts used by the researchers could lead to impurities in the final product.
More information: Ekaterina V. Vinogradova et al. Organometallic palladium reagents for cysteine bioconjugation, Nature (2015). DOI: 10.1038/nature15739
Abstract
Reactions based on transition metals have found wide use in organic synthesis, in particular for the functionalization of small molecules. However, there are very few reports of using transition-metal-based reactions to modify complex biomolecules, which is due to the need for stringent reaction conditions (for example, aqueous media, low temperature and mild pH) and the existence of multiple reactive functional groups found in biomolecules. Here we report that palladium(II) complexes can be used for efficient and highly selective cysteine conjugation (bioconjugation) reactions that are rapid and robust under a range of bio-compatible reaction conditions. The straightforward synthesis of the palladium reagents from diverse and easily accessible aryl halide and trifluoromethanesulfonate precursors makes the method highly practical, providing access to a large structural space for protein modification. The resulting aryl bioconjugates are stable towards acids, bases, oxidants and external thiol nucleophiles. The broad utility of the bioconjugation platform was further corroborated by the synthesis of new classes of stapled peptides and antibody–drug conjugates. These palladium complexes show potential as benchtop reagents for diverse bioconjugation applications.
Journal information: Nature
© 2015 Phys.org