Superbug study reveals how E. coli strain acquired deadly powers
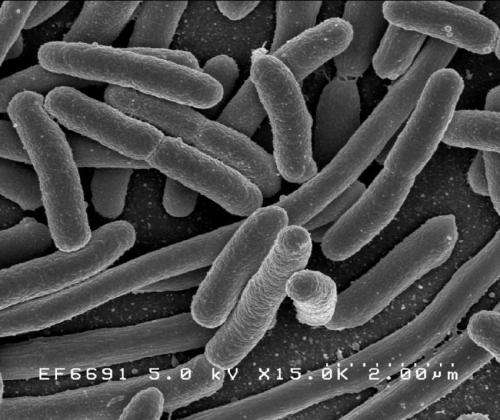
A strain of E. coli became a potentially fatal infection in the UK around 30 years ago, when it acquired a powerful toxin, a gene study has revealed.
The discovery helps to explain outbreaks of severe food poisoning that began in the 1980s.
Scientists say their findings show that E. coli O157 is continuing to evolve and should be monitored closely.
Most strains of E. coli are harmless and live in the guts of people and animals without causing illness.
E. coli O157 strains can produce molecules called shiga-toxins, which are linked to more serious human infections.
The strains that are responsible for the majority of serious illness in people in the UK produce two types of shiga-toxin - stx1 and stx2a.
A team of researchers - including scientists at Public Health England and the University of Edinburgh - decoded the genetic sequences of more than a thousand samples of E. coli O157 collected from human infections and animals over the past 30 years.
Their analysis reveals that the ancestor of E. coli O157 has been around for more than 175 years.
They found that most of the ancestor strains carry only stx1 but some strains began to acquire stx2a around 60 years ago. The dangerous strains of E. coli O157 that have caused most illness in people in the UK acquired stx2a around three decades ago, when outbreaks of severe food poisoning began to appear.
Some more recent infections are being caused by E. coli O157 strains that carry only stx2a. Early evidence suggests that these strains may be even more dangerous than those to date.
Cows are the main reservoir of E. coli O157, though they show no signs of the disease. Animals that are infected with strains that produce stx2a excrete higher levels of dangerous bacteria in their manure. This helps to spread infection between animals and increases the chances of the bacteria being passed to people.
The research was conducted by the University of Edinburgh, Public Health England, Animal Laboratories and Plant Health Agency, the Scottish E. coli Reference Laboratory, Scotland's Rural College and the University of East Anglia. Continuing work in this area is being supported by the Food Standards Agency and Food Standards Scotland. The study is published in the Microbiology Society's journal Microbial Genomics.
Professor David Gally, of the University of Edinburgh's Roslin Institute, said: "Thankfully, dangerous E. coli outbreaks remain relatively rare. Our research underlines the need to study the genetic code of strains that cause infections in humans and those present in farmed animals.
"Good hygiene practices - both with food and when out enjoying the countryside - can help to minimise the risk of these and other severe infections. Our work endeavours to understand how these toxic strains persist in cattle and the best ways to prevent them spreading to us."
Provided by University of Edinburgh