Waging war on Australia's nastiest parasite: Scientists map blowfly genome

Around 2000 genes not seen before in any other organism were discovered. These genes can now be investigated as potential drug and vaccine targets.
This blowfly is responsible for about $280 million in losses to Australia's sheep industry each year from flystrike.
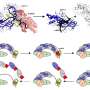
All 14,544 genes of the blowfly (Lucilia cuprina) were identified by the international research team, led by the University of Melbourne, in partnership with the Baylor College of Medicine Human Genome Sequencing Center, and funded by the United States National Human Genome Research Institute and Australian Wool Innovation.
The research, published today in Nature Communications, provides insights into the fly's molecular biology, how it interacts with the sheep's biology and, importantly, shows its potential to develop insecticide resistance.
Blowfly maggots live on the skin of sheep and invade open wounds, where they feed on tissue and cause severe skin disease, known as myiasis or flystrike. It is an aggressive and notoriously difficult pest to control.
Lead researcher on the project, Dr. Clare Anstead, of the University of Melbourne Faculty of Veterinary and Agricultural Sciences, said the genome map has 'limitless potential' for fighting the blowfly at home and abroad.
"Lucilia is a beautiful name, but it is an extremely nasty parasite. The sheep is literally eaten alive. It's horrific. The Lucilia species are responsible for more than 90 percent of flystrike in Australia and New Zealand," Anstead said.
"This fly is especially good at evolving to resist insecticides. There has been a massive amount of research into prevention and control of flystrike, from developing a vaccine, new insecticides, to targeting weak areas of the fly, and even biological control with bacteria and fungi. But none are completely effective.
"It's exciting that we have now identified more than 2000 genes that have never been seen in any other animal or plant. Some of these 'orphan' genes hold the key to the parasitic relationship between the blowfly and the sheep. They could be targeted to develop a completely new method of control."

University of Melbourne Professor Robin Gasser, who oversaw the research, added: "If you want to develop effective interventions against this fly, you need to know it inside out and understand its biology, starting by identifying all the genes. And, we have done that."
Insecticides can be effective, however, the blowflies rapidly evolve to develop resistance to these chemicals.
Professor Phil Batterham, at the University of Melbourne School of Biosciences, says this work now enables us to predict gene mutation in flies that could make them resistant to chemicals, which means we may be able to avoid the type of crisis that the medical community now faces with antibiotic resistance in bacteria.
"The next step is to isolate the parasite's 'Achilles' heel'—genes that allow the parasitic interaction between the maggots and the sheep," Batterham said.
"A vaccine that targets this gene could stop flystrike in its earliest stages. This vaccine could access vital proteins in the maggots, which would kill them. Alternatively, genomic-guided drug discovery means we could develop insecticides that selectively kill fly maggots but do not harm the host animal."
To decode the genome, researchers used a combination of supercomputing and bioinformatic techniques to handle huge reams of data.
They aim to use a powerful new technology called CRISPR to investigate switching off a number of genes, including the gene responsible for the blowfly's extraordinary sense of smell.
"Flies have an extremely sophisticated sense of smell. They can smell the difference between sheep that are resistant to the fly and those that aren't," Batterham said. "We want to produce a fly that cannot smell, so that we can understand how important that sense of smell is in the initiation of flystrike."
Lucilia cuprina is one of 30 insect species to have genome sequences generated at the Baylor College of Medicine Human Genome Sequencing Centre as part of a pilot project for the genome analysis of some 5000 arthropod species of medical, scientific, economic and agricultural importance.
Journal information: Nature Communications
Provided by University of Melbourne