May 12, 2015 report
Experimental studies of ionic interactions near a hydrophobic surface in an aqueous environment
(Phys.org)—The way a protein folds in aqueous solutions is largely determined by hydrophobic effects with the hydrophobic portions of the protein residing within the protein core or within the active site. Oftentimes, ionic interactions will occur in the hydrophobic active site. In theory, an ion pair will form a stronger interaction the closer the ions are to a hydrophobic surface, which is attributed to how the ions and the hydrophobic surface interact with the bulk water.
Even though this theory is used to describe surface interactions, there is little by way of experimental studies of ionic interactions at a hydrophobic surface. A collaborative study by groups at the University of Tokyo, the National Institute for Materials Science, University of York, Nanjing University, and the RIKEN Center for Emergent Matter science have designed an experimental model system to study ion pairing near a hydrophobic surface in an aqueous solvent. Their work appears in Science.
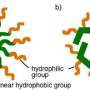
In this study, the authors created a model system in the laboratory using self-assembled monolayers (SAMs,) comprised of alkanes, to simulate a hydrophobic surface. They incorporated into their monolayer a tethered molecule with a polyethylene glycol backbone and ionic head-group. They built several monolayer systems with varying lengths of surrounding alkane chains. Each system had the same polyethylene glycol backbone and ionic head group, but different lengths of surrounding alkane chains. This allowed the authors to investigate the relationship between distance from the hydrophobic surface and ionic bonding strength.
The first two systems that they compared involved SAMs with either ethane groups or dodecane groups as the hydrophobic filler. Both SAMs systems included tetraethylene glycol with FITC, an anionic fluorescent reporter, as the ionic head group. Monolayers were formed on a silicon electrode.
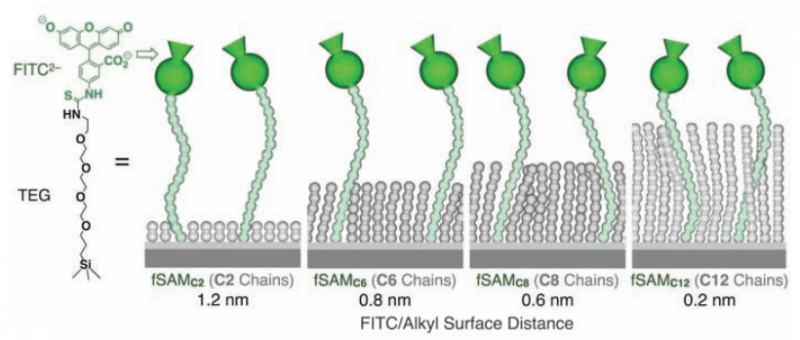
Characterization studies showed that the FITC ionic head group was 1.2 nm above the SAMs with ethane fillers and 0.2 nm above the dodecane-filled SAMs. FITC has a known fluorescence emission at 513 nm, so the first step was to test changes in FITC's fluorescence signal when it is 1.2 nm away from the hydrophobic surface and when it is 0.2 nm away. Fluorescence studies showed a similar fluorescence profile for both systems.
Both SAMs had similar titration curves, indicating a similar response to pH changes, although their fluorescence spectra were different at different pH. FITC has carboxylate groups that will protonate at lower pH and deprotonate at higher pH. At lower pH, the SAMs with ethane fillers showed a weaker fluorescence signal than the SAMs with dodecane fillers. This is likely due to conformational changes that can occur in the ethane-filled SAMs but do not occur in the dodecane SAMs due to steric hindrance. Similar results were seen when a potential from -1.5 V to 1.5 V was applied to the SAMs. This provided further confirmation that the differences were likely due to differences in steric hindrance between ethane and dodecane SAMs.
FITC can bind to guanidinium ions (Gu+) to form salt bridges. When bound to Gu+, FITC's fluorescence signal diminishes. In order to elucidate in situ ionic behavior, three Gu+ were tethered to rhodamine (Rho), a fluorophore with a strong emission band at 584 nm, to form a Gdn-Rho complex.
The next test was to see what happens when a potential is applied to each of the SAM systems when bound to Gdn-Rho. Fluorescence studies (FRET) of the ethane-filled SAMs showed that when protonation occurred, the Gdn-Rho complex was released and FITC's characteristic fluorescence emission band increased. Upon deprotonation, FITC paired with the Gdn-Rho complex, causing FITC's fluorescence signal to decrease and rhodamine's fluorescent signal to increase.
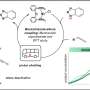
The same study, when applied to the dodecane-filled SAMs, showed different results. While the Gdn-Rho complex did bind to FITC, it was not released when a negative potential was applied to the system, and there was no change in the fluorescence signal. Titration studies of the dodecane-filled SAM showed that Gdn-Rho was released from FITC at a much lower pH compared to ethane. Calculations showed that the stability of the salt bridge increases when the distance from the hydrophobic surface decreases.
Finally, to determine how distance from a hydrophobic surface affected ionic strength between FITC and Gdn-Rho, the authors made SAMs with n-hexane and n-octane fillers. pH studies showed that the binding strength increased non-linearly as distance from the hydrophobic surface decreased. This trend is likely due to an enhanced hydrogen bonding network of the aqueous solvent near the hydrophobic surface. The ethane-filled SAMs mimicked an ionic interaction in bulk water, while the dodecane-filled SAMs mimicked an ionic interaction in water that has a stronger hydrogen-bonding network because it is near a hydrophobic surface.
These studies provide experimental evidence for theoretical measurements of ionic interactions near a hydrophobic surface.
More information: "Subnanoscale hydrophobic modulation of salt bridges in aqueous media" Science, DOI: 10.1126/science.aaa7532
ABSTRACT
Polar interactions such as electrostatic forces and hydrogen bonds play an essential role in biological molecular recognition. On a protein surface, polar interactions occur mostly in a hydrophobic environment because nonpolar amino acid residues cover ~75% of the protein surface. We report that ionic interactions on a hydrophobic surface are modulated by their subnanoscale distance to the surface. We developed a series of ionic head groups—appended self-assembled monolayers with C2, C6, C8, and C12 space-filling alkyl chains, which capture a dendritic guest via the formation of multiple salt bridges. The guest release upon protonolysis is progressively suppressed when its distance from the background hydrophobe changes from 1.2 (C2) to 0.2 (C12) nanometers, with an increase in salt bridge strength of ~3.9 kilocalories per mole.
Journal information: Science
© 2015 Phys.org