DNA of bacteria crucial to ecosystem defies explanation
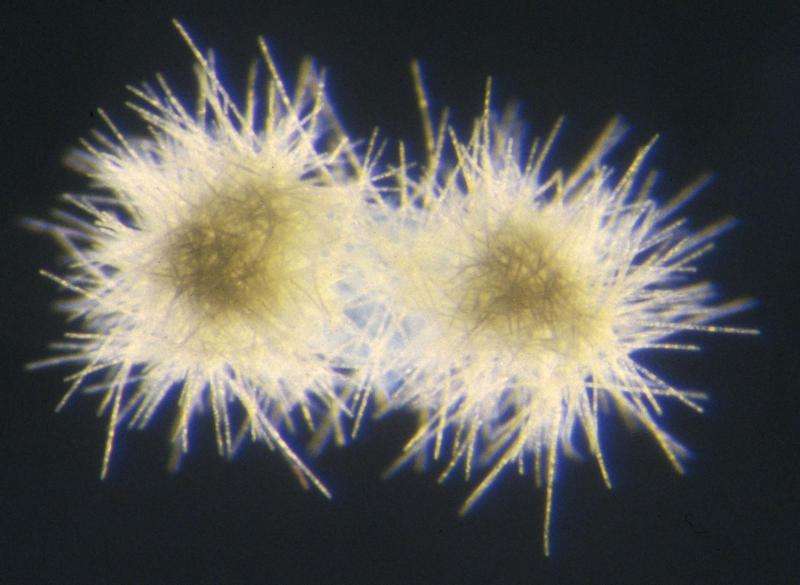
Scientists have found something they can't quite explain in one of the most barren environments on Earth: a bacterium whose DNA sequence contains elements usually only found in a much higher organism.
Trichodesmium is a type of bacteria known as an oligotroph, meaning that it can survive in incredibly nutrient-poor regions of the ocean. In fact, it thrives there—to the point that great blooms of the microorganism can be seen both with the naked eye and from satellites in space, earning it the name "sea sawdust" from ancient mariners.
This is because Trichodesmium is a "nitrogen fixer"—it makes harsh environments more habitable by turning nitrogen gas from the air into ammonium, a nutrient that other organisms can use. It's foundational to the entire food web of the ocean and therefore an important organism for scientists to understand.
By sequencing multiple Trichodesmium genomes—and using a wide variety of samples to ensure that there was no error—researchers found that only about 63 percent of the bacteria's genome is expressed as protein. That's an incredibly low amount for a bacterium and unheard of for a free-living oligotroph.
"Our study adds another wrinkle to this enigmatic organism's story," said Eric Webb, associate professor at the USC Dornsife College of Letters, Arts and Sciences and corresponding author of a study on the finding.
In a paper published in March in the Proceedings of the National Academy of Sciences, Webb and his colleagues revealed that Trichodesmium's DNA defies common evolutionary dogma, meaning that there's either an important piece of the puzzle still missing or that the understanding of the evolution of microbial genomes needs to be revisited.
"The unique evolutionary path reflected in this genome contradicts nearly all accounts of free-living microbial genome architectures to date," said lead author Nathan Walworth, a Ph.D. candidate at USC. "Different evolutionary paths are foundational to all arenas of biology, including biotechnology, so it is important for the field to be cognizant of different paths a living organism can take to achieve ecological success."
When scientists first started sequencing genomes in the last century, they found that not every part of the DNA strand encoded a protein to be expressed. In fact, we now know that only about 2 percent of the human genome is expressed—the rest was initially called "junk DNA."
Scientists now describe it as "non-coding DNA," a nod to the fact that functions have been defined for many of these regions; however, there is still controversy on the extent and role of these regions.
Despite the uncertainty, higher organisms—like humans—with tiny populations are highly vulnerable to sweeping mutations and thus can contain a lot of non-coding DNA.
Lower organisms with huge populations—like bacteria—are far less susceptible to sweeping mutations and accordingly have been shown to have genomes that are about 85 percent protein coding.
Oligotrophs, in particular, shun non-coding DNA, possibly because of the high energy-cost of living in a harsh environment. Cells need every ounce of energy simply to replicate and survive.
Trichodesmium, as Webb and his team discovered, breaks the mold. Yet despite the novelty of the finding, the ultimate cause of the large non-coding space remains to be defined.
The team theorizes that the high level of non-coding DNA is likely the result of a combination of factors, possibly including the way that Trichodesmium blooms.
Blooms make the species more vulnerable to genetic drift and can lead to genomes with enriched non-coding DNA.
"However, since there are many other bloom-forming cyanobacteria that that do not have expanded non-coding space, blooming ecology is likely not the whole story," Webb said. "Right now, we speculate that interactions with other undefined organisms might also be important."
The surprising and odd nature of Trichodesmium's DNA represents a new and open issue in the study of genetics and one, given the organism's overall importance, that the researchers are eager to answer.
More information: Trichodesmium genome maintains abundant, widespread noncoding DNA in situ, despite oligotrophic lifestyle, Nathan Walworth, 4251–4256, DOI: 10.1073/pnas.1422332112
Journal information: Proceedings of the National Academy of Sciences
Provided by University of Southern California