Molecular ruler sets bacterial needle length
When a salmonella bacterium attacks a cell, it uses a nanoscopic needle to inject it with proteins to aid the infection. If the needle is too short, the cell won't be infected. Too long, and the needle breaks. Now, University of Utah biologists report how a disposable molecular ruler or tape measure determines the length of the bacterial needle so it is just right.
The findings have potential long-term applications for developing new antibiotics against salmonella and certain other disease-causing bacteria, for designing bacteria that could inject cancer cells with chemotherapy drugs, and for helping people how to design machines at the nanoscopic or molecular scale.
The study by University of Utah biology professor Kelly Hughes and doctoral student Daniel Wee is set for online publication the week of March 16 in the journal Proceedings of the National Academy of Sciences.
"If you look at important pathogens - the bubonic plague bacterium, salmonella, shigella and plant pathogens like fire blight - they all use hypodermic-like needles to inject proteins that facilitate disease processes," Hughes says.
"Our work says that there is one mechanism - the molecular ruler - to explain how the lengths are controlled for needles in gram-negative bacteria and for hooks on flagella [the U-joints in propellers bacteria use to move] in all bacteria," he adds.
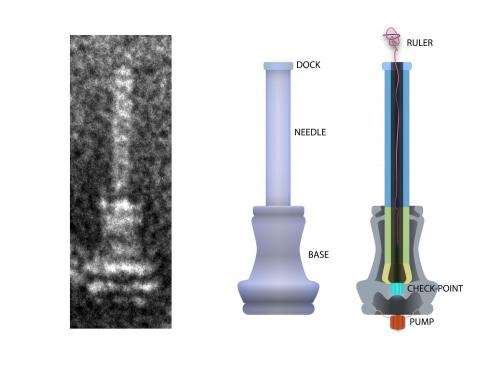
In their study, Wee and Hughes found that as a bacterial needle or "injectisome" grows, a molecular ruler - really, more like a gooey tape measure - is secreted from within the needle's base. It oozes up through the tube-like needle, and when the bottom end of the ruler reaches the bottom end of the needle, the needle stops growing and begins to inject proteins into the target cell to help the infection process.
The biologists say the National Institutes of Health-funded study refutes other theories for how salmonella and some other disease bacteria determine needle lengths.
Why the findings matter
"What we understand from bacteria can help us build nanomachines and nanobots," Hughes says, noting that bacterial flagella - the nanoscopic motor-and-propeller system they use to swim to dinner or to targets - are "the most sophisticated nanomachines in the universe."
In one example, Swiss scientists are using the design of bacterial flagella as the basis for a nanobot that will be put inside the eye to do nanoscale surgery, he adds.
In addition to flagella, a number of disease-causing bacteria also have injectisomes, which also are built of proteins, as are most structures in living organisms.
"In the case of the needle, you have a structure that extends from the surface of the bacterium like a hypodermic," Hughes says. "These needles are fragile. If one is too long, it will break off and be useless. If you make it too short, then it can't get past the surface proteins on cells it needs to invade."
By understanding how bacteria determine the length for their needles, it someday may be possible to engineer bacteria to inject chemotherapy drugs right into cancer cells.
"People would like to design bacteria that can get to cancer cells and inject poisons into just those cells and kill them, and not harm the rest of us," Hughes says.
And by understanding how certain disease-causing bacteria build their injectisomes, new antibiotics might be developed in a decade or so to target and destroy the needles and thus deter bacterial infections. The rulers that help build flagella also might be attacked by drugs to prevent bacteria from reaching target cells, "so you can kill two birds with one stone by hitting the two machines at the same time," Hughes says.
He says that approach might work against injectisome-equipped bacteria such as salmonella species that cause typhoid fever and food poisoning; shigella species that cause dysentery; the bubonic plague bacterium Yersinia pestis; disease-causing E. coli; sexually transmitted Chlamydia trachomatis; many plant pathogens; and Pseudomonas aeruginosa, which often infects burn patients and the lungs of cystic fibrosis patients.
Bacteria secrete a molecular ruler to measure needle length
Bacterial injectisomes are incredibly small, measuring only 20 to 100 nanometers long. A nanometer is one billionth of a meter, and a meter is about 39 inches long. The width of a typical human hair often is given as 100 microns, so the maximum length of a bacterial needle, 100 nanometers, is one-thousandth of the width of a human hair.
Gram-negative, disease-causing bacteria "are very closely related species, so how do they subtly control the various needle lengths to be perfect?" Hughes asks. "In one case it might be 40 nanometers versus 55 nanometers. These are small sizes. So to do this, the bacteria developed molecular rulers to differentiate needles of different lengths."
(Gram-negative bacteria are those with membranes lining both the inside and outside of their cell wall, while gram-positive bacteria have only an inner membrane.)
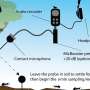
Like any cell, a bacterium is encased in a cell wall. So bacteria developed all kinds of secretions to make contact with and infect other cells: flagellar propellers to swim to food or target cells, docking structures to help bacteria stick to targets, and injectisomes to inject infection-promoting proteins into targets.
When a bacterium builds a needle, it first builds a base. "A series of proteins form a doughnut, and inside the doughnut hole, the actual secretion machine gets constructed," Hughes says. "It's the same for the flagella as it is for these needles."
Next, proteins start assembling to form the needle or injectisome.
The new study demonstrated that in salmonella, the ruler or tape measure is secreted slowly through the channel of the growing needle. Once amino acids at the bottom end of the ruler pass through the base of the needle, they tell the bacterium that the needle is long enough and to stop growing. They also tell the needle to injecting virulence proteins into the target cell, and the molecular ruler is ejected, Wee says.
How the study was performed
The new study used the Typhimurium strain of Salmonella enterica, which causes food poisoning. The researchers proved the molecular ruler determines needle length in salmonella by inserting amino acids from the plague bacterium's molecular ruler genes into genes for salmonella's molecular ruler, making rulers with seven different lengths.
Genetically engineered salmonella with seven ruler lengths were grown in a flask, their needles isolated, and the needle lengths measured under an electron microscope.
Wee found the ruler lengths correlated precisely with the lengths of the resulting needles or injectisomes, with each amino acid added to the ruler gene making the resulting needle 0.2 nanometers longer.
Previous studies found the molecular ruler determines the length of the hook or U-joint that helps turn flagella or propellers in many bacteria. Research also found the molecular ruler determines the length of both the flagellar hook and the needle in plague bacteria. But some researchers argued salmonella needle's length was determined by some other mechanism:
- One theory holds that a molecular measuring cup in the needle's base sends a cupful of needle components to assemble the needle, and the length of the needle is determined by the size of the cup. The new study disproved that by genetically removing the cup and showing that the injectisomes or needles still grew to correct lengths.
- Another theory says that as needle components assemble outside the needle's base, a rod-shaped structure assembles inside the base to link the base and needle, and that when the rod is complete, needle assembly stops, thus determining needle length. But the Utah study found the rod and needle components are not made simultaneously, but compete with each other, so as more rod parts are made, fewer needle parts are made, giving an illusion that rod completion controls needle length.
More information: Molecular ruler determines needle length for the Salmonella Spi-1 injectisome, PNAS, www.pnas.org/cgi/doi/10.1073/pnas.1423492112
Journal information: Proceedings of the National Academy of Sciences
Provided by University of Utah