A shiny, new graph query system
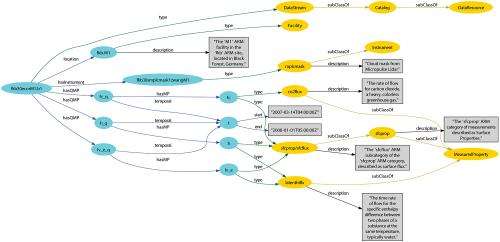
As computing tools and expertise used in conducting scientific research continue to expand, so have the enormity and diversity of the data being collected. Developed at Pacific Northwest National Laboratory, the Graph Engine for Multithreaded Systems, or GEMS, is a multilayer software system for semantic graph databases. In their work, scientists from PNNL and NVIDIA Research examined how GEMS answered queries on science metadata and compared its scaling performance against generated benchmark data sets. They showed that GEMS could answer queries over science metadata in seconds and scaled well to larger quantities of data. They also demonstrated that GEMS generally outperformed a custom-hardware solution, showing the feasibility of using cheaper, commodity hardware to obtain comparable performance.
Data standards that allow researchers to find, share, and combine information easily are becoming more essential to discover and analyze increasingly large and heterogeneous data sets. While the Semantic Web introduced the graph-based data model of the Resource Description Framework (RDF) as a way to overcome data heterogeneity, it exacerbates data volume challenges. GEMS offers a data-scalable, graph-oriented query system that has been shown to answer actual science-based queries over large-scale, real-world, curated science metadata.
As a graph engine for large clusters, GEMS works with the RDF data model—adopted in some scientific research communities—to query voluminous data sets. Currently, workable RDF databases have been shown to be 1 to 10 billion RDF triples (graph edges) in size, at which point system performance is degraded. However, by converting native science metadata to RDF triples, the data can be queried using SPARQL, the standard RDF query language. The GEMS compiler translates SPARQL queries into C++ code, which when compiled and executed, runs the query to quickly and naturally support parallel graph walking, the fundamental operation for graph queries. The Semantic Graph Library, or SGLIB, layer supports query answering, while PNNL's custom Global Memory and Threading (GMT) runtime system for clusters, at the base of the GEMS' stack, is designed to tolerate the distributed, random data access that occurs when operating on distributed graphs.
"GEMS is a distributed, in-memory, semantic graph database designed for bigger data, deeper analytics, and cheaper hardware," explained Jesse Weaver, a research computer scientist with the Analysis and Algorithms team in PNNL's Data Sciences group and the paper's primary author. "It is designed to scale out on clusters, allowing us to effectively increase global memory by adding nodes to the cluster. In our examination, GEMS was able to answer actual research project queries over science metadata in the form of 1.4 billion RDF triples on the order of seconds—a good start as we continue to tackle the problem of ever-growing volumes of data."
Ongoing GEMS development is aimed at enhancing the SPARQL-to-C++ compiler. In addition, the team is pursuing design changes that will enable queries on data sets of over 100 billion triples. GEMS' performance also will be compared with other cluster-based solutions.
More information: Weaver J, VG Castellana, A Morari, A Tumeo, S Purohit, A Chappell, D Haglin, O Villa, S Choudhury, K Schuchardt, and J Feo. 2014. "Toward a Data Scalable Solution for Facilitating Discovery of Science Resources." Parallel Computing. Early Online, September 16, 2014. DOI: 10.1016/j.parco.2014.08.002.
Provided by Pacific Northwest National Laboratory