Molecular mechanisms underlying the prevention of autoimmunity by Roquin revealed
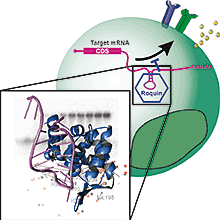
Scientists at the Helmholtz Zentrum München, the Ludwig-Maximilians University of Munich (LMU) and the Technische Universität München (TUM) have moved an important step closer to understanding molecular mechanisms of autoimmune diseases. They solved the three-dimensional structure of the Roquin protein when bound to messenger ribonucleic acid (mRNA) molecules. The results revealed that there is a much wider range of functionally important Roquin binding partners than previously assumed. The novel findings are published in the journal Nature Structural & Molecular Biology.
The Roquin protein, discovered in 2005, controls T-cell activation and differentiation by regulating the expression of certain mRNAs. In doing so, it helps to guarantee immunological tolerance and prevents immune responses against the body's own structures that can lead to autoimmune disease. Roquin is thus an immune regulator.Autoimmune diseases affect between five and ten per cent of the population. They usually occur as a result of complex environmental influences when a genetic predisposition exists. Only in rare cases the development of the disease is determined by a single mutated gene. However, a single mutation in the Roquin gene in a mouse model was shown to be responsible for the development of the autoimmune disease systemic lupus erythematosus. This mutation in the Roquin protein also led to a high susceptibility to type 1 diabetes and rheumatoid arthritis and induced angioimmunoblastic T-cell lymphoma.
Elucidation of the three-dimensional structure of the Roquin-RNA complex
An interdisciplinary team comprising the research groups led by Prof. Michael Sattler, Dr. Dierk Niessing and Prof. Vigo Heissmeyer at the Helmholtz Zentrum München, Ludwig-Maximilian University (LMU) and the Technische Universität München (TUM) has now revealed unprecedented insight into how Roquin recognizes its RNA binding partner and thereby controls T-cell functions. To this end, the scientists Dr. Andreas Schlundt, Gitta Heinz, and Dr. Robert Janowski used the X-ray crystallography platform of the Helmholtz Zentrum München to determine the spatial structure of the RNA binding domain of Roquin when bound to its RNA target. The interaction of Roquin with additional RNA binding partners was studied in solution using nuclear magnetic resonance (NMR) spectroscopy at the Bavarian NMR Center, a joint research infrastructure of the Helmholtz Zentrum München and TUM. Furthermore, the researchers could confirm the biological significance of the molecular recognition of the RNA by studying Roquin-dependent gene regulation in cellular systems.
The results obtained reveal for the first time the molecular interactions with which roquin recognizes a binding motif in a gene's mRNA. "To our surprise, these results indicate that a greater range of binding modes plays an important functional role for the gene regulation in T-cells," says Prof. Michael Sattler. "Thus, our findings suggest that Roquin regulates a larger number of genes than was previously assumed," Dr. Niessing adds. In addition to the mRNAs with optimal recognition motifs, which are tightly bound and predominantly regulated by Roquin, there is a potentially much larger number of mRNAs which are more weakly bound, but nevertheless regulated by Roquin. "On the basis of these findings we will now focus on understanding how Roquin levels are regulated in T-cells, since strong and weakly bound target mRNAs will experience a principally different regulation when the availability of the protein varies" explains Prof. Vigo Heissmeyer.
Basis for developing treatment
Defining the molecular interplay between Roquin and RNA is a prerequisite for con-trolling the function of Roquin and using its role for therapeutic strategies to treat autoimmune diseases. To this end, the scientists are now planning follow-up studies to find out how the function of Roquin can be manipulated.
More information: Schlundt A. et al. (2014). "Structural basis for RNA recognition in roquin-mediated post-transcriptional gene regulation," Nature Structural & Molecular Biology (2014) DOI: 10.1038/nsmb.2855. Received 14 April 2014 Accepted 03 June 2014 Published online 13 July 2014
Journal information: Nature Structural & Molecular Biology
Provided by Helmholtz Association of German Research Centres