MRI, on a molecular scale: Team develops system that could peer into atomic structure of individual molecules

For decades, scientists have used techniques such as X-ray crystallography and nuclear magnetic resonance (NMR) imaging to gain invaluable insight into the atomic structure of molecules. Such efforts have long been hampered by the fact that they demand large quantities of a specific molecule, often in ordered and crystalized form, to be effective—making it all but impossible to peer into the structure of most molecules.
Harvard researchers say those problems may soon be a thing of the past.
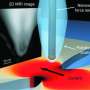
A team of scientists, led by Professor of Physics and of Applied Physics Amir Yacoby, has developed a magnetic resonance imaging (MRI) system that can produce nanoscale images, and may one day allow researchers to peer into the atomic structure of individual molecules. Their work is described in a March 23 paper in Nature Nanotechnology.
"What we've demonstrated in this new paper is the ability to get very high spatial resolution, and a fully operational MRI technology," Yacoby said. "This work is directed toward obtaining detailed information on molecular structure. If we can image a single molecule and identify that there is a hydrogen atom here and a carbon there … we can obtain information about the structure of many molecules that cannot be imaged by any other technique today."
Though not yet precise enough to capture atomic-scale images of a single molecule, the system already has been used to capture images of single electron spins. As the system is refined, Yacoby said he expects it eventually will be precise enough to peer into the structure of molecules.
While the system designed by Yacoby and colleagues operates in much the same way conventional MRIs do, the similarities end there.
"What we've done, essentially, is to take a conventional MRI and miniaturize it," Yacoby said. "Functionally, it operates in the same way, but in doing that, we've had to change some of the components, and that has enabled us to achieve far greater resolution than conventional systems."

Yacoby said that while conventional systems can achieve resolutions of less than a millimeter, they are effectively limited by the magnetic field gradient they can produce. Since those gradients fade dramatically within just feet, conventional systems built around massive magnets are designed to create a field large enough to image an object—like a human—that may be a meter or more in length.
The nanoscale system devised by Yacoby and colleagues, by comparison, uses a magnet that's just 20 nanometers in diameter—about 300 times smaller than a red blood cell—but is able to generate a magnetic field gradient 100,000 times larger than even the most powerful conventional systems.
The difference, Yacoby explained, is that the nanoscale magnet can be brought incredibly close, within a few billionths of a meter, to the object being imaged.
"By doing that, we can achieve spatial resolution that's far better than one nanometer," he said.
The departures from conventional MRI systems, however, didn't end there.
To construct a sensor that could read how molecules react to that magnetic field gradient, Yacoby and colleagues turned to a field that would appear to be unconnected to imaging—quantum computing.
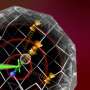
Using ultra-pure, lab-grown diamonds, the team milled tiny devices, each of which ended in a super-fine tip, and embedded an atomic-scale impurity, called a nitrogen-vacancy (NV) center in each tip, creating a single quantum bit, or qubit—the essential building block of all quantum computers.
In experiments published last year, Yacoby and his collaborators showed that as the tip was scanned across the surface of a diamond crystal, the quantum bit interacted with electron spins near the crystal's surface. Those interactions could then be used to create an image of individual electron spins. However, while the sensitivity of the quantum bit sensor is sufficient to detect individual electron spins and represents a quantum leap forward from earlier efforts, its spatial resolution is limited by its distance from the object that is being imaged.
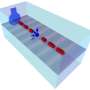
To create truly 3-D images, Yacoby and colleagues combined the quantum-bit sensing approach with the large-field gradient by bringing the nanomagnet in close proximity to both the sample of interest and the qubit sensor. By scanning the magnet in 3-D, but very close to the sample, they were able to detect individual electron spins as they reacted to the magnetic field.
"This is really a game of bringing both the magnet very close to generate large gradients, and bringing the detector very close to get larger signals," Yacoby said. "It's that combination that gives us both the spatial resolution and the detectability.
"Our current system is already capable of imaging individual electron spins with sub-nm [subnanometer] resolution," he said. "The goal, eventually, is to put a molecule in proximity to our NV center to try to see the components within that molecule, namely the nuclear spins of the individual atoms composing it. This is by no means an easy task, since the nuclear spin generates a signal that is 1,000 times smaller than that of the electron spin … but that's where we're headed."
More information: "Nanoscale MRI: Dark spins in the spotlight." Lloyd Hollenberg. Nature Nanotechnology 9, 253–255 (2014). DOI: 10.1038/nnano.2014.65. Published online 23 March 2014. dx.doi.org/10.1038/nnano.2014.65
Journal information: Nature Nanotechnology
Provided by Harvard University