New sequencing tools give up close look at yeast evolution
Using next-generation sequencing, corresponding author Gianni Liti et. al. provide a detailed characterization of the genetic variation present within the baker's yeast species.
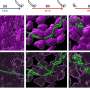
The baker's yeast Saccharomyces cerevisiae has been associated with human activities for thousands of years, being the primary biological agent in baking, brewing, winemaking and other fermentation processes. It is also one of the most important model organisms in molecular biology and genetics research. For a long time, the history and evolution of this important yeast has been a completely mystery, but recent advances in genome sequencing technologies now allow it to be studied in great detail.
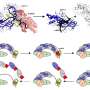
Using next-generation sequencing, corresponding author Gianni Liti et. al. provide a detailed characterization of the genetic variation present within the baker's yeast species. They sequenced the genomes of 42 strains of S. cerevisiae and its closest relative S. paradoxus, which is an entirely wild species that has not had any contact with humans. A central finding of this study is that even though strains in S. paradoxus are separated by much greater genetic distances in terms of single-nucleotide polymorphisms (SNPs), the S. cerevisiae strain genomes harbor more variation in terms of absence and presence and copy number of genes. It has previously been observed that trait variation is also much larger in S. cerevisiae than in its wild relative. These new results therefore raise the intriguing hypothesis that this variation in the content of the genome, rather than single-nucleotide differences, underlies the large phenotypic variation in S. cerevisiae.
The authors find that the subtelomeric regions of the genomes, located just before the telomeres at each chromosome end, are highly enriched for genome variation that is likely to contribute to differences in traits between strains. This includes loss-of-function mutations that likely disrupt the function of whole genes. As an example of functional variation they describe how differences in the copy number of a subtelomeric gene cluster controls the ability of strains to grow under arsenic stress, and demonstrate that this variation is the product of convergent evolution in yeast lineages in different parts of the world.
"These genome sequences allowed us to expose surprising differences between the evolutionary histories of the common baker's yeast and its wild relative. Our results suggest that the very large diversity in traits observed between strains of baker's yeast might mostly be due to the presence or absence of entire genes rather than differences in single DNA letters."
The study provides intriguing insights into the recent history of this important organism and the relationship between genome variation and trait variation. Future research will further elucidate what role humans have played in shaping the evolution of baker's yeast, for example the extent to which the genomic variation is a consequence of yeast strains moving into novel habitats and niches opened up by human activities.
The study is published in this week's Molecular Biology and Evolution.
Journal information: Molecular Biology and Evolution
Provided by Oxford University Press