Team discovers new form of virus reproduction
Each small step that Science takes to discover how viruses infect cells is always very valuable to researchers and society, since it provides relevant information to fight infections.
However, the recent discovery made by the Ikerbasque Professor Nicola Abrescia, at the Center for Cooperative Research CIC bioGUNE, in collaboration with the University of Helsinki, has been more than just a small step. For the first time, a new mechanism used by viruses to infect cells has been described. This mechanism involves the formation of a tube generated with pre-existing lipids and proteins in the structure of the virus. This study, carried out with a virus model system – bacteriophage (infects bacteria) PRD1 – leads the way forward in the fight against bacterial infections such as E. coli or Salmonella.
The breakthrough is also very important since the virus that has been analysed is representative of other viruses that infect all the kinds of existing cells: archaea, bacteria and eukaryotic. In other words, they can infect animal and plant cells, bacteria and other types of cells.
Virus with lipid vesicle
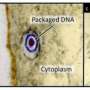
Unlike bacteria, which are classified as an independent cell domain, viruses are acellular structures that depend on their ability to infect cells and transfer their genetic material in order to ensure their survival and propagation, as they are not able to reproduce on their own. They are basically composed of a genome (DNA or RNA) and a capsid, in other words, a protein structure that envelopes and protects this genetic material. Some have a lipid vesicle enveloping the capsid and, in other cases, the vesicle is surrounded by the capsid.
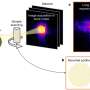
Research at the CIC bioGUNE, in collaboration with other European centres, has discovered a new viral phenomenon in which viruses with internal lipid vesicles and specifically bacteriophage PRD1, representative of this virus family, injects its genome into the cell to infect it.
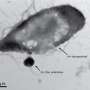
This study has earned its publication in the prestigious scientific journal, PLoS Biology, and has been included as one the most outstanding papers in the Synopsis section. The research team observed how the virus develops a proteolipid tail to create a conductive tube, penetrates the bacterial cell and transfers its genome into it in order to replicate.
Research conclusions
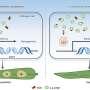
One of the main conclusions of the study is that bacteriophage PRD1 uses the proteins of the membrane in its lipid vesicle and the lipids to assemble the tube. Then it drills a hole in the bacteria, which is an essential step to infect it. Researchers also observed that the tube, one of the smallest that has been studied, only allows the passage of one double-stranded DNA molecule.
"The bacteriophage PRD1 is special in the sense that it does not have a constitutively attached tail. Instead, when it infects bacteria, it creates a tube to transfer its genetic material inside the cell. The virus manages to form the tubes by restructuring the icosahedral capsid and remodelling the internal vesicle, which is rich in membrane proteins," Abrescia declares.
Researchers were able to visualize the structure of this tube and the process of infecting bacteria, using not only powerful electron microscopes, but also advanced electron microscopy and image processing techniques available at CIC bioGUNE.
Possible antibiotics
This research is not only interesting because it is the first time that the infection process of a member of this virus family has been described, but also because PRD1, the studied virus, infects and kills bacteria such as E. coli and Salmonella. Therefore, the knowledge generated could lead towards the development of new fighting strategies against infections caused by these bacteria.
"If we understands how this virus drills and penetrates the cell that it is going to infect, we can potentially think of recreating artificial tubes in the laboratories that are similar to those generated by the virus PRD1, to be used as antibiotics," Abrescia concludes.
More information: Peralta, B. et al. Mechanism of Membranous Tunnelling Nanotube Formation in Viral Genome Delivery, PLOS. BIOL, September 2013. www.plosbiology.org/article/info%3Adoi%2F10.1371%2Fjournal.pbio.1001667
Journal information: PLoS Biology
Provided by CIC bioGUNE