Discarded data may hold the key to a sharper view of molecules
(Phys.org) -- There's nothing like a new pair of eyeglasses to bring fine details into sharp relief. For scientists who study the large molecules of life from proteins to DNA, the equivalent of new lenses have come in the form of an advanced method for analyzing data from X-ray crystallography experiments.
The findings, just published in the journal Science, could lead to new understanding of the molecules that drive processes in biology, medical diagnostics, nanotechnology and other fields.
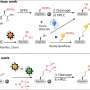
Like dentists who use X-rays to find tooth decay, scientists use X-rays to reveal the shape and structure of DNA, proteins, minerals and other molecules.
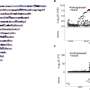
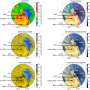
As X-rays pass through atoms they reflect distinctive patterns, which reveal what atoms are present and how atoms are bonded to each other. However, some data are typically discarded because of concerns over quality. In particular, data derived from edge regions of the pattern — although very important for understanding the details of structure — are often overwhelmed by the random errors associated with a weak signal in the midst of a lot of background noise.
Oregon State University biophysicist Andy Karplus and his colleague Kay Diederichs at the University of Konstanz in Germany have now proven that useful information can be gleaned from data that have about five times the noise level that was previously considered acceptable.
"The criteria that have been used in the past are way too conservative," said Karplus. "These data that people have been throwing out are actually good."
The bottom line for crystallographers is the accuracy of their molecular models. The better the model, the better it will predict the pattern created by X-rays passing through a molecule, and the better it will be to develop new drugs and nanotechnologies that operate at the molecular scale.
The new method may be the most important conceptual advance in the past 20 years in how these data are used in modeling, the scientists said. It shows how data from "noisy" parts of the measurement can still provide information, and allows scientists to see directly where the model is limited by noise in the data and where the model is a better estimate of molecular structure than experimental data.
"The question is, 'Where do we cut it off?'" said Karplus. By adding data at incremental steps and showing how the model improved, Karplus and Diederichs showed that scientists had been cutting off their analyses too soon and discarding data that could sharpen their view of molecular structure.
"The big impact on the field will be that every structure determined from here on out will be a little more accurate because people won't throw away data that are okay," Karplus said. "If you have a crummy image of the protein, it will get a little sharper. If you have a good image of the protein, it will also get a little sharper."
While the method will be an important step for X-ray crystallographers, the scientists said that other physical sciences may also find ways to benefit from this type of data quality analysis. They noted that one branch of science has been using this type of statistical analysis for many years. The field of psychometrics — the analysis of data from psychological tests — has used a similar technique called the "Spearman-Brown prophecy formula" to determine the minimum length of such tests.
"Now that we know that very noisy data are useful, this will presumably enable still further improvements as it stimulates new software development to do a better job of handling such weak data," said Karplus.
More information: Paper: "Linking Crystallographic Model and Data Quality," by P.A. Karplus at Oregon State University in Corvallis, OR; K. Diederichs at University of Konstanz in Konstanz, Germany.
The paper is also the subject of a Perspectives ("Resolving Some Old Problems in Protein Crystallography") piece in the same issue of Science by Phil Evans of the MRC Laboratory of Molecular Biology in Cambridge, England.
Journal information: Science
Provided by Oregon State University