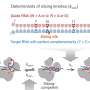
Small DNA circles found outside the chromosomes in mammalian cells and tissues, including human cells
Researchers from the University of North Carolina at Chapel Hill have helped identify a new DNA entity in mammalian cells and provided evidence that their generation leaves behind deletions in different locations of the cells’ genetic program, or genome.
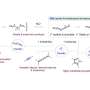
The researchers discovered these new entities in mouse tissues and brain cells and in human cell lines. Unlike previously identified large DNA circles, these circles of unique non-repetitive microDNA sequences are in the coding and control regions of genetic information.
The study was published online March 8, 2012, in the journal Science. It was led by University of Virginia authors Yoshiuyuki Shibata, PhD, senior research associate, Pankaj Kumar, PhD, bioinformatician, and Anindya Dutta, MD, PhD, Byrd Professor and Chair of Biochemistry and Molecular Genetics.
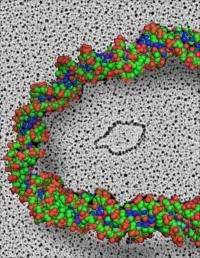
UNC researchers Jack D. Griffith, PhD, Kenan Distinguised Professor of microbiology and immunology and a member of the UNC Lineberger Cancer Center, and Smaranda Willcox, research analyst, performed electron microscopic analysis that provided the visual evidence of the microDNA sequences.
“Jack Griffith’s group are the world’s experts on electron microscopy of nucleic acids, so naturally we turned to them to see if we could visualize the microDNAs that our molecular biology experiments were pointing to,” said Dutta. “Seeing is believing.”
The known DNA in cells are in nuclear chromosomes that are millions of base pair long linear stretches of DNA capped by telomeres, like the plastic tips of shoelaces. MicroDNAs are 200-400 base pair long circles in the nucleus that are not attached to chromosomes, making them new DNA entities.
According to Dutta, their result is surprising in that it indicates that occasionally DNA replication is sloppy. Despite proof-reading activity and repair mechanisms, sometimes the copying process eliminates little snippets of DNA as circles and leaves behind microdeletions in the chromosomes.
Thus, there is some element of luck in what the daughter cells in tissues are getting in terms of genetic material. For example, some cells in the hippocampus portion of the brain may have a small deletion on one copy of a gene A, while another set of cells in the same tissue may have a small deletion in one copy of another gene B. Often these microdeletions are silent, meaning they do not affect gene expression. However, by random chance, they can sometimes be in critical areas where they affect a cell’s function. Thus, there is a possibility that all cells in a given tissue have slightly different DNA.
“Smaranda Willcox’s images also revealed that some circles have only one DNA strand instead of the usual two, which adds an unexpected twist to an already novel story,” said Griffith.
While there is not yet proof that the microdeletions actually cause disease, the very fact that these disparities exist in the genome, or genetic program delivered to individual cells, means that simply by chance some cells may have a non-functional or low functioning gene.
Normally, every cell in the human body has two copies of a gene – one each from mother and father. However, if one copy has a pre-existing mutation and the other copy has a microdeletion, the result could be problematic. The researchers theorize that future work in this area could lead to new knowledge about the causes of autism or schizophrenia, which may be due to incorrect functioning of certain genes in brain tissue.
Microdeletions in protective genes, such as tumor suppressors, could render them inactive and thus reduce protections against cancer, so this discovery is relevant to cancer research.
“This is a basic science discovery to explain the general mechanism of DNA loss, which can have applications for further research that may lead to new knowledge about specific health conditions later on,” said UNC’s Willcox who carried out the EM study.