A 'wild card' in your genes
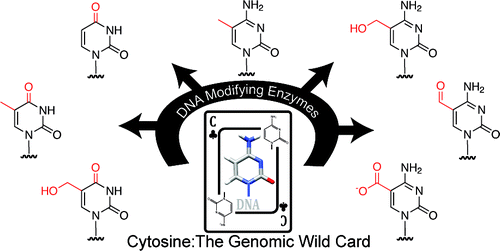
The human genome and the endowments of genes in other animals and plants are like a deck of poker cards containing a "wild card" that in a genetic sense introduces an element of variety and surprise that has a key role in life. That's what scientists are describing in a review of more than 100 studies on the topic that appears in ACS Chemical Biology.
Rahul Kohli and colleagues focus on cytosine, one of the four chemical "bases" that comprise the alphabet that the genetic material DNA uses to spell out everything from hair and eye color to risk of certain diseases. But far from just storing information, cytosine has acquired a number of other functions that give it a claim to being the genome's wild card. "In poker, the rules of the game can occasionally change," they note in the article. "Adding a 'wild card' to the mix introduces a new degree of variety and presents opportunities for a skilled player to steal the pot. Given that evolution is governed by the same principles of risk and reward that are common to a poker game, it is perhaps not surprising that a genomic 'wild card' has an integral role in biology."
They discuss the many faces of cytosine that make it such a game-changer and the biological processes that help to change its identity. Removing something called an amine group from cytosine, for instance, allows the immune system to recognize and destroy foreign invaders such as viruses. Adding so-called "methyl groups" on cytosines acts as on/off switches for genes. The authors say that these many faces of cytosine allow it to play various roles and give it true "wild card" status.
More information: The Curious Chemical Biology of Cytosine: Deamination, Methylation,and Oxidation as Modulators of Genomic Potential, ACS Chem. Biol., Article ASAP. DOI: 10.1021/cb2002895
Abstract
A multitude of functions have evolved around cytosine within DNA, endowing the base with physiological significance beyond simple information storage. This versatility arises from enzymes that chemically modify cytosine to expand the potential of the genome. Some modifications alter coding sequences, such as deamination of cytosine by AID/APOBEC enzymes to generate immunologic or virologic diversity. Other modifications are critical to epigenetic control, altering gene expression or cellular identity. Of these, cytosine methylation is well understood, in contrast to recently discovered modifications, such as oxidation by TET enzymes to 5-hydroxymethylcytosine. Further complexity results from cytosine demethylation, an enigmatic process that impacts cellular pluripotency. Recent insights help us to propose an integrated DNA demethylation model, accounting for contributions from cytosine oxidation, deamination, and base excision repair. Taken together, this rich medley of alterations renders cytosine a genomic “wild card”, whose context-dependent functions make the base far more than a static letter in the code of life.
Provided by American Chemical Society
















