Digital microfluidics opening the way for revolution in blood sampling
The days of the blood sample routine - arm out, tie tube, make a fist, find a vein and tap in -- may soon be over, thanks to a new analysis method developed at U of T by Institute of Biomaterials and Biomedical Engineering (IBBME) core professor Aaron Wheelerin which only a pinprick of blood is necessary.
Traditional methods of blood sampling requires intravenous extraction of several millilitres of blood. A phlebotomist then separates serum, which is frozen for transport or storage and later thawed and analyzed. A relatively new alternative to the traditional method uses blood samples stored as dried blood spots (DBSs).
The DBS method requires only a pinprick to extract a few microlitres of blood, which is blotted onto filter paper, where the sample, it has been found, remains stable. While DBSs have been gaining increasing popularity for the ease of sampling and storage for some time, they are still not a standard laboratory technique, and the process for using them remained laborious -- until now.
In a study published in Lab on a Chip last week, Wheeler and colleagues demonstrated the proof-of-principle that digital microfluidics could be used to automate the process of dried blood spot analysis in the case of testing for specific genetic diseases at Newborn Screening Ontario (NSO) in Ottawa. This paper is the result of a collaboration between Wheeler and NSO rsearchers.
NSO regularly screens every baby born in Ontario for genetic diseases - some 140 000 babies a year - and collects DBS samples via heelprick. Each DBS must be manually collected. Technicians must prepare the sample for testing, put it into a centrifugal tube, pipette solvent onto the sample, extract the necessary material by centrifuge, and then use robotics to conduct the chemical analysis.
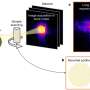
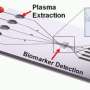
Wheeler’s digital microfluidic platform automates this process. Droplets are manipulated onto the sample using electrical signals, and the material needed for analysis is extracted - all on a “lab-on-a-chip” with little manual intervention. Wheeler, the Canada Research Chair in Bioanalytical Chemistry, created the prototype for this process in the Bahen Cleanroom, a facility of the Emerging Communications Technology Institute at U of T.
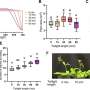
Wheeler’s study quantified particular amino acids that are markers of three metabolic disorders: phenylketonuria, homocystinuria, and tyrosinemia. His next steps will be to evaluate the rest of the 28 diseases that NSO screens for.
Wheeler’s innovation is indicative of the innovative tools for biomedical engineering that IBBME researchers create. “The applications for this process go far beyond newborn screening,” Wheeler stated. “Pharmaceutical companies are moving towards dried blood spot analysis, but they’re still lacking the tools to make widespread use feasible. We’ve demonstrated that digital microfluidics could be that tool. Our system is fast, robust, precise, and compatible with automation.”
While it might be a while before the days of the dreaded blood sample needle are behind us, Wheeler’s digital microfluidics method is the next step in moving to a DBS-based sampling system, said Pranesh Chakraborty, director of NSO. “This approach could save considerable costs as a result of the lower volumes of reagent required,” he affirmed. “An automated system based on this approach would also process samples faster, with higher accuracy, less risk of errors, all while freeing up time for technologists to perform other work.” Charaborty’s team provided the screening and medical perspective in this research.
A patent has been filed, and Wheeler, who also holds appointments in chemistry and Banting and Best Department of Medical Research, is currently exploring commercialization options.
Provided by University of Toronto