Into the (mis)fold: a diagnostic tool for proteins
(PhysOrg.com) -- Alzheimer’s disease is the most common form of dementia, currently affecting more than 35 million people worldwide. Although many genetic and hereditary factors are thought to contribute to the telltale deterioration of memory and cognitive functions resulting from Alzheimer’s, a central aspect to this disease is an accumulation of misfolded proteins in the brain.
Now, scientists at Berkeley Lab have engineered a universal, highly sensitive technique for detecting misfolded proteins in biological fluids. This groundbreaking nanoscience capability could help pinpoint Alzheimer’s in its early stages and enable researchers to discover new therapies for this devastating disease.
When a protein doesn’t fold into its normal shape, it also doesn’t perform its normal functions. This disruption in behavior could lead to proteins that aggregate into plaques or deposits and become toxic to cells. In Alzheimer’s disease, aggregates of a protein called beta-amyloid form in the central nervous system, causing damage to cells in the brain and triggering dementia.
An analytical capability for measuring tiny clusters of these proteins—before irreversible damage occurs—would be a powerful tool in the early detection of Alzheimer’s and other misfolded protein diseases. However, despite significant research efforts, there are currently no diagnostic tools available to selectively detect small-scale aggregates of misfolded proteins in biological fluids, such as blood or spinal fluid.
“This collaboration illustrates how a biomedical problem can also be a nanoscience problem, in which a chemical reagent is needed to recognize partially aggregated proteins,” said Ron Zuckermann, Director of the Biological Nanostructures Facility at the Molecular Foundry, a nanoscience user facility at Berkeley Lab. “We were faced with the challenge of synthesizing a material that’s capable of specifically detecting this aggregated protein and not any of the other proteins in the blood.”
Zuckermann is a pioneer in the development of peptoids, synthetic polymers that behave like naturally occurring proteins but can withstand aggressive chemical and biological environments without degrading. His group previously discovered peptoids capable of self-assembling into nanoscale jaws, nanosheets, and nanoscale ropes that braid themselves.
“Peptoids are ideal for this application as they are similar to proteins in structure, but different enough that they aren’t degraded by enzymes in the blood,” added Zuckermann. “We can now engineer materials that are capable of specific recognition yet can evade destruction.”
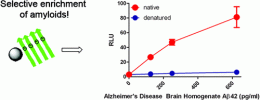
Using the Foundry’s state-of-the-art robotic synthesis capabilities, the team prepared a panel of peptoids designed to capture a misfolded prion protein, an abnormal, infectious form of a cellular protein found in the brain. By attaching these peptoids to tiny magnetic beads, the team could then use a magnet to isolate misfolded proteins directly from blood samples. The most selective and sensitive of these peptoids, coined aggregate-specific reagent, or ASR1, could capture not only the prion aggregates, but aggregates associated with Alzheimer’s disease as well.
“Our study shows how basic research capabilities can be translated into a practical application,” said Zuckermann. “The potential for this tool to serve as a diagnostic in other misfolded protein diseases, such as Parkinson’s and Type II diabetes, is wide open, and I’m excited to continue this collaboration.”
More information: This research is reported in a paper titled, “A universal method for detection of amyloidogenic misfolded proteins,” appearing in the journal Biochemistry and available in Biochemistry online.
Abstract
Diseases associated with the misfolding of endogenous proteins, such as Alzheimer’s disease and type II diabetes, are becoming increasingly prevalent. The pathophysiology of these diseases is not totally understood, but mounting evidence suggests that the misfolded protein aggregates themselves may be toxic to cells and serve as key mediators of cell death. As such, an assay that can detect aggregates in a sensitive and selective fashion could provide the basis for early detection of disease, before cellular damage occurs. Here we report the evolution of a reagent that can selectively capture diverse misfolded proteins by interacting with a common supramolecular feature of protein aggregates. By coupling this enrichment tool with protein specific immunoassays, diverse misfolded proteins and sub-femtomole amounts of oligomeric aggregates can be detected in complex biological matrices. We anticipate that this near-universal approach for quantitative misfolded protein detection will become a useful research tool for better understanding amyloidogenic protein pathology as well as serve as the basis for early detection of misfolded protein diseases.
Provided by Lawrence Berkeley National Laboratory


















