Key step identified in Legionnaire's disease infection process
(PhysOrg.com) -- NIH researchers have uncovered a key step in the biochemical sequence the bacterium which causes Legionnaire’s disease uses to reproduce inside the cells it infects.
The bacterium is known to activate a cell protein to help it hide from the cell’s defenses while it reproduces. In the current study, the researchers have discovered how the bacterium switches off the protein so that its offspring can leave the cell and begin the infection process anew.
The finding may one day lead to new ways to treat Legionnaire’s disease and diseases caused by related bacteria.
Legionnaires’ disease is a form of pneumonia caused by several types of Legionella bacteria. The disease was named after its discovery in a group of people attending an American Legion convention in 1976. The bacteria live in fresh water and are often found in hot tubs, air conditioning cooling units, and fountains. The disease occurs in individuals of all ages, including infants. It primarily affects people with weakened or compromised immune systems, including those who are over 65 years of age, those with lung disease, and smokers. The disease is spread when people inhale droplets of water containing the bacteria.
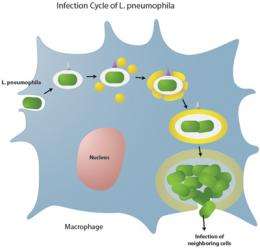
The infection process begins when Legionella bacteria are engulfed by immune cells known as macrophages, explained the study’s senior author, Matthias P. Machner, Ph.D., head of the Unit on Microbial Pathogenesis at the NIH’s Eunice Kennedy Shriver National Institute of Child Health and Human Development. Macrophages patrol the lungs for potential disease-causing organisms and envelop them in balloon-like membrane structures known as phagosomes. Ordinarily, foreign organisms are held inside the phagosome until they are destroyed by chemical reactions that take place inside the cell.
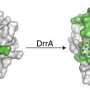
Previous studies conducted by Dr. Machner and other researchers have shown that the Legionella bacteria hide out inside the phagosomes, subverting the cell’s defense machinery for its own purposes. The bacterium forces the cell to camouflage the phagosome. The bacterium does this by hijacking Rab1, a protein that the cell makes. Enzymes the bacterium produces chemically change Rab1, adding to it a molecule known as AMP (adenosine monophosphate). The altered Rab1 rests on the surface of the phagosome. There, it serves as a kind of beacon, attracting cell structures known as transport vesicles. The vesicles are bubble-like structures ordinarily used for ferrying proteins throughout the cell. The vesicles eventually cover the surface of the phagosome and fuse with its membrane, thereby hiding it from the cell’s defenses.
"Adding AMP to Rab1 is like jamming a switch so that it’s permanently in the ‘on’ position," Dr. Machner said. "This way the bacterium commands the host cell to continuously deliver vesicles to the phagosome until the disguise is completed."
Hidden inside the camouflaged phagosome, the bacterium then reproduces. Eventually, the bacterium destroys the host cell membrane, opening up pores through which the new bacteria leave the macrophage. Once outside the cell, the new bacteria infect other macrophages, and the cycle repeats itself.
In the current study, Dr. Machner and his colleagues sought to find out how the Legionella bacterium can turn off the jammed switch on Rab1, thereby inactivating the beacon that attracts vesicles to the Legionella-containing phagosome.
The researchers searched the bacteria’s genes and found the information needed to produce a protein called SidD. In a series of experiments, the researchers found that the SidD protein sliced the AMP molecule from Rab1, which stopped summoning vesicles to the surface of the phagosome.
"SidD switches off Rab1 when the bacterium no longer needs it," Dr. Machner said.
Dr. Machner explained that the finding ultimately may lead to new approaches for treating Legionella infections. The finding may also have implications for treating other diseases as well. In addition to Legionella, other species of disease-causing bacteria also use the strategy of adding AMP to host cell proteins. These include Vibrio parahemolyticus, which can contaminate seafood and cause food poisoning, and Haemophilus somni, which can cause a blood infection in cattle.
Provided by National Institutes of Health