'On the origin of nematodes' -- A phylogenetic tree of the world’s most numerous group of animals
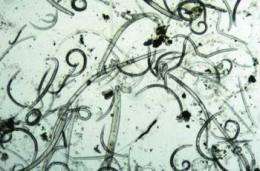
Scientists from Wageningen University and Research Centre have published the largest nematode Phylogenetic Tree to date in cooperation with the Dutch Plant Protection Service (PD) and the University of California (Riverside) in the November issue of the journal Nematology.
"It contains over 1,200 species and is entirely based on the analysis of DNA sequence data. It is relatively straightforward - and in fact we've shown it already - to define species-specific DNA barcodes on the basis of this data set that allows for the detection of nematodes in soil with an unprecedented accuracy," scientists said.
Nematodes are the world’s most numerous group of animals with two to 20 million individuals, usually smaller than one millimetre, per square metre of soil. These nematodes include a minority that can cause diseases to humans, animals or plants. Unfortunately these pathogenic organisms share a strong resemblance to each other as well as to useful nematode species. This makes finding out which nematodes are present in a soil of a given area an extremely time-consuming and a specialist task.
The international group of scientists studying nematode DNA selected a specific element of the DNA that codes for a major part of ribosomes, parts of the cells of both plants and animals responsible for the production of proteins. Containing 1,700 building blocks, this piece of DNA allowed scientists to distinguish between most nematode species. In fact the resolution of the datya set appeared to bre much better than we had ever expected.
Based on the DNA-analyses, the scientists could make some major steps forward towards to the reconstruction of the evolution of this successful group of animals, including the ones that - because of their feeding behaviour - cause major damage to lifestock and crops. Our results provided sufficient information for distinguishing a number of plant-parasitic nematodes. The new technology has already been used to study tens of thousands of soil samples. Faster and more accurate than traditional microscopic analyses, the technology has been an immediate success.
More information: Hanny Van Megen, Sven van den Elsen, Martijn Holterman, Gerrit Karssen, Paul Mooyman, Tom Bongers, Oleksandr Holovachov, Jaap Bakker and Johannes Helder (2009). A Phylogenetic Tree of Nematodes based on about 1200 full-length Small Subunit Ribosomal DNA sequences. Nematology Vol. 11(6), 927-950
Provided by Wageningen University