New windows opened on cell-to-cell interactions (w/ Video)

Applying biological molecules from cell membranes to the surfaces of artificial materials is opening peepholes on the very basics of cell-to-cell interaction.
Two recently published papers by a University of Oregon biophysicist and colleagues suggest that putting lipids and other cell membrane components on manufactured surfaces could lead to new classes of self-assembling materials for use in precision optics, nanotechnology, electronics and pharmaceuticals.
Though the findings are basic, they provide new directions for research to help understand nature at nanotechnological scales where the orientation of minuscule proteins is crucial, said Raghuveer Parthasarathy, who is a member of the UO's Material Science Institute, the Institute of Molecular Biology and the Oregon Nanoscience and Microtechnologies Institute (ONAMI).
Controlling molecular orientation from cell membranes
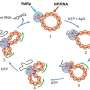
In a paper appearing online in the Journal of the American Chemical Society (JACS) in early July, Parthasarathy teamed with organic chemists at the University of California, Berkeley, to study how molecules are oriented on their cell membranes to allow for cell-to-cell interactions.
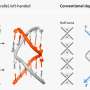
The six-member research team built tiny artificial molecules that mimic brush-like membrane proteins and contain tiny fluorescent probes at the outer end. These miniscule polymers were incorporated into artificial membranes placed on a silicon wafer that acts like a mirror, allowing precise optical measurements of the orientation of the molecule.
Electron microscopy revealed the presence of rigid, rod-like brushy glycoprotein (sugar-containing compounds) -- 30 billionths of a meter long -- similar to natural cell-surface proteins. Interaction between cells occurs when these rods stand up from the membranes, a property whose control remains poorly understood.
The surprise, Parthasarathy said, was that the sugar-laden rods stood up like trees rising in a forest only for particular fluorescent probes, which represented just 2 percent of the molecule's weight.
The big issue that surfaced from the project -- funded by the U.S. Department of Energy, National Science Foundation and the Alfred P. Sloan Foundation -- was that the slightest trepidation of a molecule's structure affects its orientation, he said.
The goal, Parthasarathy said, may be to determine how to control the orientation of the brush-like forest through either chemical or optical measures to, in turn, control cell interaction. Such control of artificially produced molecules, he added, could have huge potential applications in the electronics industry.
Parthasarathy's UO team is now looking at DNA anchored to membranes to compare the findings and see if such on-off switching of the orientation of molecules may be possible.
"There are brush-like proteins at cell surfaces that are really important for such things as cellular interactions within the immune system," Parthasarathy said. "At the surface of every cell is a forest of molecules to induce interactions. These proteins need to rise from the forest. What allows them to stick up or lie down? We've really had a poor idea of what's going on. Knowing the genome and what proteins are there is crucially important, but that information in itself does not tell you anything about the answer to the question."
Source: University of Oregon (news : web)