Researchers identify proteins making up mechanosensitive ion channels
Researchers at Washington University in St. Louis are the first to identify two proteins responsible for mechanosensitive ion channel activities in plant roots. Scientists have long known that plant cells respond to physical forces. Until now, however, the proteins controlling the ion channel response remained a mystery.
As the name suggests, mechanosensitive channels are paths through the cell membrane that respond to mechanical forces such as gravity, pressure, or touch. Under certain forces, a channel opens, allowing the flow of ions, such as calcium and potassium ions, into and out of the cell. Different forces might close the channel, stopping the flow.
This cross-membrane ion flow has been measured electrophysically, using a technique called the patch-clamp method. But the molecular nature of the channels themselves was not known. Now, knowing the proteins involved makes it possible to discover what the channels do for the whole plant.
"People have been characterizing mechanosensitive channels in plants for 20 years," said Elizabeth Haswell, Ph.D., assistant professor of biology at Washington University in St. Louis and lead investigator of this project, "This is the first time anybody has been able to show which proteins underlie these activities."
Plants do it, bacteria do it
The two proteins governing ion channels in Arabidopsis root are MSL9 and MSL10, according to the study published in the May 20 issue of Current Biology. MSL stands for MscS-Like proteins because of their similarity to a family of bacterial channels known as MscS (mechanosensitive channels of small conductance). Even though bacteria and plants are not closely related in terms of evolution, this study shows that bacterial and plant cells are probably using the same types of proteins to respond to mechanical forces.
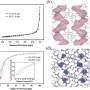
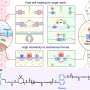
To establish that the channels were in fact mechanosensitive, Haswell's French colleagues used the patch-clamp method to measure the movement of ions across the membrane of Arabidopsis root cells as the pressure inside the cell increased. These experiments demonstrated that increasing cellular pressure also increased the ion flow across the membrane. Likewise, as the pressure inside the cell went down, the ion flow decreased.
To determine whether MSL9 and MSL10 were responsible for this ion flow, Haswell created a mutant line of Arabidopsis without either type of protein. When the root cells of the plants lacking MSL9 and MSL10 were tested, the researchers saw very little change in ion flow across the membrane as the pressure inside the cell increased. In other words, without these two proteins, very little channel activity was seen. And the little channel activity they did measure was shown to be caused by different proteins.
Two to tango
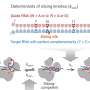
Having shown that MSL9 and MSL10 were responsible for the ion channel activity, Haswell and colleagues set out to determine if both were required for the response or if only one did most of the work. Therefore, they tested plants that lacked only one of each protein and were surprised to discover that cells with only MSL9 showed one type of activity and cells with only MSL10 showed a different type of activity. And, importantly, cells with both proteins showed a third type, suggesting that both MSL9 and MSL10 are required to produce the mechanosensitive channel activity seen in wild-type Arabidopsis root.
Haswell and colleagues propose that the channel is composed of subunits of both proteins MSL9 and MSL10 and that this combined structure results in the unique mechanosensitive ion channel behavior observed in the wild-type plants and not in any of the mutant lines.
Despite identifying proteins that govern this ion channel response in Arabidopsis root, mysteries remain. To determine how the mutant plants might be defective and reveal the purpose of the channels, Haswell's group grew the plants that lacked these channels under challenging conditions, including high salt, root barriers, and dehydration. "We tried hundreds of experiments, but we never saw a difference between the mutant and wild-type," said Haswell, "but that's definitely one of the next big steps – to find out what the channels really do and why they're important."
Source: Washington University in St. Louis