RNA enzyme structure offers a glimpse into the origins of life
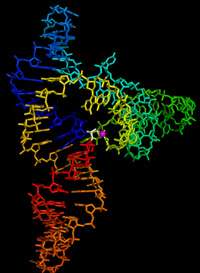
Researchers at the University of California, Santa Cruz, have determined the three-dimensional structure of an RNA enzyme, or "ribozyme," that carries out a fundamental reaction required to make new RNA molecules. Their results provide insight into what may have been the first self-replicating molecule to arise billions of years ago on the evolutionary path toward the emergence of life.
In all forms of life known today, the synthesis of DNA and RNA molecules is carried out by enzymes made of proteins. The instructions for making those proteins are contained in genes made of DNA or RNA (nucleic acids). The circularity of this process poses a challenge for theories about the origins of life.
"Which came first, nucleic acids or proteins? This question once seemed an intractable paradox, but with the discovery of ribozymes, it is now possible to imagine a prebiotic 'RNA World' in which self-replicating ribozymes accomplished both tasks," said William Scott, associate professor of chemistry and biochemistry at UC Santa Cruz.
Scott and postdoctoral researcher Michael Robertson determined the structure of a ribozyme that joins two RNA subunits together in the same reaction that is carried out in biological systems by the protein known as RNA polymerase. Their findings are published in the March 16 issue of the journal Science.
"An RNA-dependent RNA polymerase ribozyme is the foundation of the entire RNA World hypothesis," Robertson said. "With that, you would have an RNA capable of making copies of itself; mutations or errors in some copies would result in variations that would be acted on by Darwinian natural selection, and the molecules would evolve into bigger and better ribozymes. That's what makes this structure so interesting."
Robertson and Scott determined the structure of a ribozyme that is not an entirely self-replicating RNA molecule, but it does carry out the fundamental reaction required of such a molecule--a "ligase" reaction creating a bond between two RNA subunits.
Robertson obtained the ligase ribozyme through a kind of test-tube evolution when he was a graduate student at the University of Texas, Austin, working in the lab of biochemist Andrew Ellington. Starting with a mixture of randomly synthesized RNA molecules and selecting for the desired properties, researchers are able to evolve RNA enzymes from scratch. In the Ellington lab, Robertson evolved the ligase ribozyme (called the L1 ligase) and determined which parts were critical for its function and which parts could be removed to create a "minimal construct."
At UC Santa Cruz, he began trying to grow crystals of the ribozyme so that he could use x-ray crystallography to determine its structure. Crystallizing RNA molecules is extremely difficult, and Robertson tried dozens of different versions of the ribozyme under different conditions before he succeeded. Using x-ray crystallography--which involves shining a beam of x-rays through the crystals and analyzing the resulting diffraction patterns--Robertson and Scott were then able to determine the three-dimensional structure of the ribozyme.
The ribozyme has three stems that radiate from a central hub. The active site where ligation occurs is located on one stem, and the structure shows that the molecule folds in such a way that parts of another stem are positioned over the ligation site, forming a pocket where the reaction takes place. A magnesium ion bound to one stem and positioned in the pocket plays an important role in the reaction, Robertson said.
The structure indicates that this artificially selected ribozyme uses reaction mechanisms that are much like those used by naturally occuring enzymes, Robertson said.
"The L1 ligase appears to use strategies of transition-state stabilization and acid-base catalysis similar to those that exist for natural ribozymes and protein enzymes," he said.
Source: University of California - Santa Cruz