Novel optical tweezers instrument unravels bacterial DNA
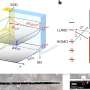
VU Amsterdam researchers have developed an optical tweezers instrument, which they used to unravel bacterial chromosomes. The researchers, headed by Dr. Gijs Wuite, have demonstrated how an important protein, called H-NS, bridges DNA strands in bacteria. Thanks to this technology, it has now been proven that the seemingly chaotic cluster of bacterial DNA is in fact organized and can function dynamically. Moreover, the H-NS protein is a potential target for developing medication to treat bacterial infections.
The research findings will be published in the scientific journal Nature on November 16, 2006.
Unlike cells in the human body, bacteria do not have a nucleus. These micro-organisms are much less complex than our human body cells, but this, rather surprisingly, makes it more difficult to determine how the DNA in a bacterial cell is organized. Prior to the use of the newly developed optical tweezers instrument, it was very difficult to study the spatial organization of bacterial DNA.
In human and animal cells, DNA-strands are coiled up inside chromosomes and extremely well organized. The bacterial chromosome is much more dynamically organized by a small group of proteins that non-specifically bind the DNA. Consequently, these proteins have more, and more general, functions. The DNA appears to be unorganized, like a ball of noodles in the cell – or so it seemed at least.
For cell division or DNA repair, the bacterium must duplicate its DNA, and this cannot be done without choreographed order. DNA duplication is the result of (among other factors) the action of DNA binding motor proteins: they slide along the DNA and replicate every nucleotide in the DNA-sequence. It was already known that certain proteins prevented the DNA from becoming entangled; but what was unknown is how it was then possible for a motor protein to slide along the DNA-strands. This mystery has now been solved.
Gijs Wuite, Remus Dame and Maarten Noom, the authors of the article to be published in Nature, began by demonstrating that a specific protein (namely, histone-like nucleoid structuring protein, H-NS) bridges two DNA strands. H-NS is a small protein that has on both its ends a small, ball-like element that can attach to DNA, probably fitting in the small cavities along the DNA’s spiral staircase-like structure. Remus Dame: “It’s great that in our measurements the helical shape of the DNA emerges. But what is much more important is that we were able to measure the strength with which the H-NS is bound to the DNA.” It is a weak bond: each H-NS arm is loosely bound to a DNA-helix.
Moreover this bond is unstable: over a certain period of time, the arm of the H-NS comes loose, in order to then reattach itself to the DNA. Because there is a lot of H-NS protein between the two parallel DNA-helices, the overall bridging activity is unhindered if each protein occasionally let’s go and then reattaches itself. Gijs Wuite: “And this precisely explains why motor proteins are unhindered by H-NS when they move along the DNA: the force these proteins exert is greater, and H-NS simply allows them to pass. This has never before been demonstrated.”
On the Net: www.nature.com/nature/journal/ … 444/n7117/index.html
Source: VU University Amsterdam