Researchers publish first genomic collection of human microbes
The Human Microbiome Project (HMP) today published an analysis of 178 genomes from microbes that live in or on the human body. The researchers discovered novel genes and proteins that serve functions in human health and disease, adding a new level of understanding to what is known about the complexity and diversity of these organisms.
The human microbiome consists of all the microorganisms that reside in or on the human body. Outnumbering cells in the human body by 10 to 1, some of the microorganisms cause illnesses, but many are necessary for good health. Currently, researchers can grow only some of the bacteria, fungi and viruses in a laboratory setting. However, new genomic techniques can identify minute amounts of microbial DNA in an individual and determine its identity by comparing the genetic signature to known sequences in the project's data base. The paper is published in the May 21 issue of the journal Science.
"This initial work lays the foundation for this ambitious project and is critical for understanding the role that the microbiome plays in human health and disease," said National Institutes of Health Director Francis S. Collins, M.D., Ph.D. "We are only at the very beginning of a fascinating voyage that will transform how we diagnose, treat and ultimately, prevent many health conditions."
Launched in 2008 as part of the NIH Common Fund's Roadmap for Medical Research, the HMP is a $157 million, five-year effort that will implement a series of increasingly complicated studies that reveal the interactive role of the microbiome in human health.
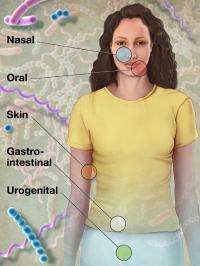
The 178 microbial genomes in this report launch the HMP reference collection that eventually will total approximately 900 microbial genomes of bacteria, viruses and fungi. These data will then be used by HMP researchers to characterize the microbial communities found in samples taken from healthy human volunteers and, later, those with specific illnesses. Samples are currently being collected for HMP from five areas of the body: the digestive tract, the mouth, the skin, the nose and the vagina.
"Although this is only the first step in making HMP medically useful, we already have learned surprising things about the diversity and complexity of the microorganisms that live in and on our body," said Jane Peterson, Ph.D., associate director of the NHGRI Division of Extramural Researcher and a leader of the HMP effort. "The next stages of this coordinated study will begin to associate the presence or absence of specific micro-organisms with various states of health and illness."
Researchers also conducted a preliminary survey to gain insights into the function of some of the newly identified genes and proteins unique to individual microbial strains. For instance, researchers found previously unknown proteins produced by bacteria that live in the stomach that may cause gastric ulceration, a hole in the stomach lining. In addition, they found a small number of newly identified novel proteins associated with how sugars and amino acids are metabolized.
Researchers also evaluated the microbial diversity present in the HMP reference collection. For example, they found 29,693 previously undiscovered, unique proteins in the reference collection - more proteins than there are estimated genes in the human genome. They compared their results to the same number of previously sequenced microbial genomes randomly selected from public databases. In the microbial genome from public databases, they found 14,064 novel proteins. These data, the researchers say, suggest that the HMP reference collection has nearly twice the amount of microbial diversity than is represented by microbial genomes already in public databases.
One of the primary goals of the HMP reference collection is to expand researchers' ability to interpret data from metagenomic studies. Metagenomics is the study of a collection of genetic material (genomes) from a mixed community of organisms. Comparing metagenomic sequence data with genomes in the reference collection can help researchers determine whether they are novel or already existing sequences.
To evaluate whether the reference collection of genomes was meeting the goal above, the researchers compared 16.8 million microbial sequences found in public databases to the genome sequences in the HMP reference collection. They found that 62 genomes in the reference collection showed similarity with 11.3 million microbial sequences in public databases and 6.9 million of these - about 41 percent - correspond with genome sequences in the reference collection.
This analysis demonstrates that genomes sequenced as part of the reference collection add directly to an understanding of the human microbiome. However, researchers cautioned that at least one-third of the metagenomic sequences are still not represented by any genome in the reference collection and that this analysis focused only on the gastrointestinal tract. The authors added that additional genomes likely exist in other body sites and the completion of the reference collection should address many of the remaining organisms not accounted for in this analysis.
The initial stage of the HMP, which includes the current study, focused on bacteria, but future genome sequencing and human microbiome studies also will capture information about more complex microbes and viruses. The effort so far also has allowed researchers to create a framework for data resources and standards. In addition, the project is supporting the development of innovative technologies and computational tools, coordination of data analysis, and an examination of some of the ethical, legal and social implications of human microbiome research.
Genome sequencing work for the project is done by the HMP-funded large-scale sequencing centers: the Human Genome Sequencing Center, Baylor College of Medicine, Houston; Washington University Genome Sequencing Center, Washington University School of Medicine, St. Louis; The J. Craig Venter Institute, Rockville, Md.; and the Broad Sequencing Platform, Broad Institute of MIT/ Harvard, Cambridge, Mass.
The HMP is currently funding pilot demonstration projects by researchers that will sample the microbiomes of healthy volunteers and volunteers with specific diseases over the next year. This will allow researchers to study changes in the microbiome at particular body sites in healthy controls compared to patients affected by diseases. These studies will use samples collected from seven areas of the body: the digestive tract, the mouth, the skin, the nose, the vagina, the blood and the male urethra.
More information:
HMP data may also be accessed from its Data Analysis and Coordination Center website, hmpdacc.org/
The Human Microbiome Jumpstart Reference Strains Consortium. A Catalog of Reference Genomes from the Human Microbiome. Science. 21 May 2010:Vol. 328. no. 5981, pp. 994 - 999. DOI:10.1126/science.1183605
Provided by National Institutes of Health















