Weird, ultra-small microbes turn up in acidic mine drainage
(PhysOrg.com) -- In the depths of a former copper mine in Northern California dwell what may be the smallest, most stripped-down forms of life ever discovered.
The microbes — members of the domain of one-celled creatures called Archaea — are smaller than other known microorganisms, rivaled in size only by a microbe that can survive solely as a parasite attached to the outside of other cells. Their genomes, reconstructed by a group at the University of California, Berkeley, are among the smallest ever reported.
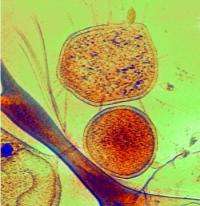
The researchers also discovered another mine-dwelling microbe that occasionally produces weird protuberances unlike any structures seen before in Archaea and uses them to penetrate the ultra-small microbes.
"Other cells in the mine have what looks like a needle that sometimes pokes right into the cells," said Brett J. Baker, a researcher in UC Berkeley's Department of Earth and Planetary Science and first author of a new paper describing the findings. "It is really remarkable and suggests an interaction that has never been described before in nature."
Baker and a team led by Jillian Banfield, UC Berkeley professor of earth and planetary science and of environmental science, policy and management and staff scientist at the Lawrence Berkeley National Laboratory (LBNL), published their findings last week in the online early edition of the journal Proceedings of the National Academy of Sciences.
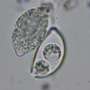
Under a light microscope, the ultra-small microbes look like specks of dust. But LBNL microscopist Luis R. Comolli used a state-of-the-art cryoelectron microscope, or cryoEM, to obtain high-resolution, 3-D images and even measure an individual microbe's internal volume - about one-hundredth the volume of an E. coli bacterium. Each of the microbes, dubbed ARMAN, for archaeal Richmond Mine acidophilic nanoorganisms, is only 300-500 nanometers in diameter, one-third the diameter of E. coli.
The team reconstructed the genomes of three distinct lineages of ARMAN and found them to be tiny — a mere 1 million base pairs, in contrast to hundreds of billions in humans. In the smallest of the three, the average gene length is only 774 base pairs, in contrast to the average gene length in humans of 10,000 to 15,000 base pairs. Base pairs, the smallest chemical units of the gene, are nucleic acids that come in four forms. The base pairs are chained together to make DNA, and a gene is a sequence of base-pairs coding for a unique protein.
The genomes are so small that the researchers initially suspected that the ARMAN microbes are parasites upon other microbes, since parasites can afford to lose genes that their host already has.
But of the 70 individual specimens so far imaged in 3-D, 90 percent seem to be free-living. The other 10 percent are impaled on the mysterious needle-like spines of Thermoplasmatales, the other Archaea living alongside ARMAN in the mine. The researchers suspect that the penetrating spines may mean that the microbes live off other microbes at least part of the time, unlike symbiotic organisms or parasites, which must always associate with other organisms to live.
"ARMAN are among the smallest microbes we know of that, if not free-living, are at least not permanently obliged to be a parasite or symbiont," Comolli said.
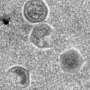
The cells are about as large as the largest viruses, which can replicate only in living organisms and are not considered to be "living."
"The genome is very compact," Baker added." A microbial genome 10 percent larger has the same number of genes as ARMAN."
The organism has a much higher percentage — 45 percent — of unknown genes than any other organism sequenced, he said.
"ARMAN share a lot of genes with Euryarchaeota and Crenarchaeaota, but they also have a lot of genes not seen before in these branches of Archaea," he said, suggesting that ARMAN may have been around since these two branches split billions of years ago.
The cryoEM tomographic images show that ARMAN, like other Archaea, have no nucleus, but they do have an enigmatic internal tube.
Banfield's group first described the ARMAN microbes four years ago, after identifying the organisms in acidic pools in the Richmond Mine, which is owned by Ted Arman, in Iron Mountain, Calif. The team's continued analysis has revealed amazing organization within the mine drainage biofilm communities that grow on solutions with the acidity of battery acid. The new data will help the researchers explore even further the community of organisms in the mine and determine how the organisms are able to live in such harsh environs and convert iron sulfides to sulfuric acid.
"Having these microbes described at the genomic level allows us to develop molecular identification methods and combine these methods with a 3-D view of the microbes to study the distribution of these organisms within this little ecological system, this little society, in the mine," Comolli said.
Provided by University of California - Berkeley