The Nobel Prize and Pond Scum as a 'Model' Organism
A man is a man and a mouse is a mouse, but if you talk to a few biomedical scientists about their research, at least one is likely to spring the term “mouse model” on you.
But there actually are many so-called model organisms. They are used as stand-ins to learn more about what most of us are really interested in - ourselves, our health, our afflictions and how our bodies work. Cost, convenience and how accurately a critter can model particular biological processes shared with humans play a role in the choice of organism employed. The mouse often comes out on top.
But UCSF’s Elizabeth Blackburn, PhD, will receive a share of this year’s Nobel Prize for Physiology or Medicine for studies she began in the 1970s on a model organism that has just one cell. When Blackburn began the prize-winning work, she was looking to solve a puzzle about DNA and cell division that that was so fundamental, it pertained to protozoans as well as people.
Blackburn sought to discover how cells can overcome a biochemical step during cell division that results in a shortening of the ends of DNA molecules each time a cell replicates its genetic material.
Without something to protect the DNA, one would think that humans would be unable to develop normally, and that protozoans would go extinct as successive generations lost another bit of the genetic blueprint.
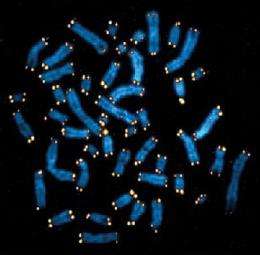
It turns out that the DNA in each chromosome has expendable, protective DNA capping it, so that gene-encoding DNA is not lost during cell division. Blackburn was the first to describe this DNA, called a telomere. Later in her own lab, Blackburn worked with Carol Greider, then a graduate student, to identify an enzyme called telomerase, which can lengthen and replenish telomeres in each chromosome within the cell. Telomere length and telomerase now are known to play a part in cancer and in other human diseases.
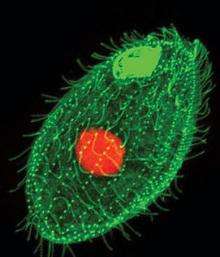
But why did Blackburn start with Tetrahymena, a single-celled protozoan she often refers to as “pond scum”? Why not study human cells or cells or creature more similar to us?
For that, we turn to Joseph Gall, PhD, now a researcher at the Carnegie Institution for Science, Baltimore, and an adjunct professor at Johns Hopkins University. It was in Gall’s lab that Blackburn first spelled out the sequence of DNA building blocks that define the telomere of Tetrahymena.
“She did the original work on telomeres in my lab at Yale as a postdoc. The origination of the field of telomeres is due to the use of the protozoan Tetrahymena as an experimental organism.
“Every chromosome has only two ends. From a biochemical standpoint, with the techniques available in the 1970s, it would have been totally impossible to define those ends, because they would have accounted for too small a fraction of the total human DNA. But in Tetrahymena the genome is broken up into thousands of pieces, and so there are thousands of ends. Therefore it was possible for Liz to concentrate on one particular type of molecule and to define the ends. So, the organism is very important.
“She had been a graduate student in Fred Sanger’s lab at Cambridge in England, and from there she brought a new technique of DNA sequencing. Nobody in the lab had done DNA sequencing before. With the combination of the sequencing technique and the material [Tetrahymena DNA] she was able to sequence the ends of the molecules, and that turned out to be the Tetrahymena telomere sequence.
“Later in the studies she did with Carol Greider, Tetrahymena was absolutely essential, because it’s one of the few organisms that make enough of the enzyme that puts these ends on chromosomes, which made it possible to study the enzyme in a biochemical fashion. It was the choice of the organism and the material that was absolutely critical.”
As for the Nobel committee’s decision to give the prize to Blackburn, along with Greider, PhD, now on the faculty of Johns Hopkins University, and Jack Szostak, PhD, from Massachusetts General Hospital, Gall says, “It totally appropriate, it’s really great that it recognizes this important work.”
Provided by UCSF