Scientists overcome nanotech hurdle
When you make a new material on a nanoscale how can you see what you have made? A team lead by a Biotechnology and Biological Sciences research Council (BBSRC) fellow has made a significant step toward overcoming this major challenge faced by nanotechnology scientists.
With new research published today (13 August) in ChemBioChem, the team from the University of Liverpool, The School of Pharmacy (University of London) and the University of Leeds, show that they have developed a technique to examine tiny protein molecules called peptides on the surface of a gold nanoparticle. This is the first time scientists have been able to build a detailed picture of self-assembled peptides on a nanoparticle and it offers the promise of new ways to design and manufacture novel materials on the tiniest scale - one of the key aims of nanoscience.
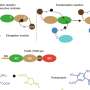
Engineering new materials through assembly of complex, but tiny, components is difficult for scientists. However, nature has become adept at engineering nanoscale building blocks, e.g. proteins and RNA. These are able to form dynamic and efficient nanomachines such as the cell's protein assembly machine (the ribosome) and minute motors used for swimming by bacteria. The BBSRC-funded team, led by Dr Raphaël Lévy, has borrowed from nature, developing a way of constructing complex nanoscale building blocks through initiating self-assembly of peptides on the surface of a metal nanoparticle. Whilst this approach can provide a massive number and diversity of new materials relatively easily, the challenge is to be able to examine the structure of the material.
Using a chemistry-based approach and computer modelling, Dr Lévy has been able to measure the distance between the peptides where they sit assembled on the gold nanoparticle. The technique exploits the ability to distinguish between two types of connection or 'cross-link' - one that joins different parts of the same molecule (intramolecular), and another that joins together two separate molecules (intermolecular). As two peptides get closer together there is a transition between the two different types of connection.
Computer simulations allow the scientists to measure the distance at which this transition occurs, and therefore to apply it as a sort of molecular ruler. Information obtained through this combination of chemistry and computer molecular dynamics shows that the interactions between peptides leads to a nanoparticle that is relatively organized, but not uniform. This is the first time it has been possible to measure distances between peptides on a nanoparticle and the first time computer simulations have been used to model a single layer of self-assembled peptides.
Dr Lévy said: "As nanotechnology scientists we face a challenge similar to the one faced by structural biologists half a century ago: determining the structure with atomic scale precision of a whole range of nanoscale materials. By using a combination of chemistry and computer simulation we have been able to demonstrate a method by which we can start to see what is going on at the nanoscale.
"If we can understand how peptides self-assemble at the surface of a nanoparticle, we can open up a route towards the design and synthesis of nanoparticles that have complex surfaces. These particles could find applications in the biomedical sciences, for example to deliver drugs to a particular target in the body, or to design sensitive diagnostic tests. In the longer term, these particles could also find applications in new generations of electronic components."
Professor Nigel Brown, BBSRC Director of Science and Technology, said: "Bionanotechnology holds great promise for the future. We may be able to create stronger, lighter and more durable materials, or new medical applications. Basic science and techniques for working at the nanoscale are providing the understanding that will permit future such applications of bionanotechnology."
Source: BBSRC