Scientists Automate Molecular Evolution
Under the control of a computer at The Scripps Research Institute, a population of billions of genes morphed through 500 cycles of forced adaptation to emerge as molecules that could grow faster and faster on a continually dwindling source of chemical fuel -- a feat that researchers describe as an example of "Darwinian evolution on a chip."
The super molecules that resulted, a species of RNA enzyme, were produced in about 70 hours using an automated tool that is about the size of a compact disc, according to the study published in the April issue of PLoS Biology. The Scripps Research investigators who designed the device note that the findings provide an example of the Darwinian principle of selective pressure at work, seen in real time.
"This is evolution at the level of molecules as a fact, not a theory," says the study's senior investigator, Gerald Joyce, Scripps Research professor in the Departments of Chemistry and Molecular Biology. "This is what it looks like when a computer controls conditions that push molecules to adapt in order to thrive--survival of the fittest on the smallest scale possible."
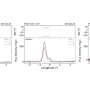
The evolved enzymes that resulted exhibited a new set of 11 mutations that improved their ability to survive under substrate-starvation conditions by 90-fold, compared to the starting molecules.
The study's first author, Brian Paegel, a postdoctoral researcher in Joyce's lab, noted that the study is the first of its kind. In previous research, scientists had managed to force adaptation in the test tube by manually adding and extracting ingredients. However, that technique produced an isolated snapshot rather than a dynamic overview of the evolution process.
"No one has been able to observe what the process looks like until now," Paegel says. "It's like before you could only see little bits of a fine painting. Now, we can step back and watch a complete picture of evolution happening at its most fundamental level, on a molecular scale."
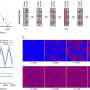
Paegel and Joyce designed and patented the microfluidic device they used in the study. The device is basically a thin glass plate, four inches in diameter, with microscopic channels and valves that a computer can control to add or extract small amounts of material. The cost to construct a device is about $8.
Pushing the Limits of an Artificial Molecule
In this study, the scientists used artificial RNA enzymes based on molecules originally developed by David Bartel and Jack Szostak at Harvard Medical School, which were derived starting from completely random RNA sequences. These molecules had been used before in "test tube" evolution experiments carried out by manual methods. The molecules have the ability to catalyze the joining of other RNA molecules, similar to a large protein known as an RNA polymerase. In the system developed by Paegel and Joyce, an RNA molecule that performed this reaction would automatically be copied to produce molecular "progeny."
In the newly published study, Paegel and Joyce loaded the microfluidic device with billions of RNA enzymes and the RNA copying machinery. They then added the chemical fuel that the RNA enzymes must utilize in order to be copied. The scientists provided progressively lower concentrations of the fuel at set intervals, as a way to direct the evolution of the RNA enzymes. Every time the concentration was reduced, those RNA enzymes whose genetic features allowed them to withstand the more stringent conditions multiplied in greater numbers than RNA enzymes that were not so adapted. Each time the size of the population of molecules reached a predetermined level, the computer isolated 1/10 th of the population--which now contained higher numbers of successfully adapted RNA enzymes—and mixed it with a new supply of chemical fuel.
These steps were repeated automatically for 500 iterations of 10-fold growth followed by 10-fold dilution. "The competition between the RNA enzymes to scrape up the few substrates became progressively stiffer, and the variants of RNA enzymes that could bind fastest and tightest to the substrate fuel molecules won out," Paegel says.
"We starved these enzymes, pushing them to become better and faster at forming a bond so they could reproduce themselves," Joyce says. "This is like the evolution of animals that can survive food famines. Only here we can see it happen in 70 hours and we know why the mutations that constitute evolution in these molecules occurred. We witnessed the entire story."
Although RNA molecules are not "alive" in the classic sense, they evolve in the same way that viruses do, Paegel says. But unlike those pathogens, which need protective casings to survive, these molecules are not dangerous, he says, because they degrade quickly outside of the chip.
Besides offering a powerful demonstration of real-time evolution—which could also be used to study adaptation in proteins, viruses, and even cellular organisms--the technology may have a number of practical uses, the scientists say, although it was not designed with these in mind. For example, it may be possible to use the technology to create RNA enzymes that act as super chemical sensors. The technology might also be used to help in the design of new medicines by encouraging molecules to evolve to perform a desired function.
The study was funded by grants from the National Science Foundation, the National Aeronautics and Space Administration, and the National Institutes of Health. To view the article, go to biology.plosjournals.org/perls … document&doi=10.1371%2Fjournal.pbio.0060085 .
Source: Scripps Research Institute