Evolution in the nanoworld

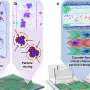
The automatic molecular assembly and selection steps exhibited by the molecules, which start as random mixtures, demonstrates a fundamental step in the evolution of life. The organization is activated by instructions which are built-in to the molecules. During assembly, molecules exhibit active selection: those in incorrect positions move to make room for others which fit properly.
The molecular-level observation of such self-selection gives, for the first time, direct insight into fundamental steps of the biological evolution from inanimate molecules to living entities. The resulting nanostructures also hold great promise as an efficient avenue to new catalysts, nanotechnologies, and surface applications.
In the Proceedings of the National Academy of Sciences, the scientists from the research groups of Klaus Kern at the Max Planck Institute for Solid State Research in Stuttgart (MPI) and of Mario Ruben at the Karlsruhe Institute of Technology (KIT) explain that this observation of molecular organization at surfaces may lead to further insight of how simple, inanimate molecules can build up biological entities of increasing structural and functional complexity, such as membranes, cells, leaves, trees, etc.
"The ability of molecules to selectively sort themselves in highly organized structures is a fundamental requirement for all molecular based systems, including biological organisms," explains Prof. Dr. Klaus Kern, director of the Nanoscale Science Department at the MPI.
Dr. Mario Ruben's research team at KIT is responsible for designing molecules with built-in instructions, which when read out activate the self-selection process. He comments: "Spontaneous ordering from random mixtures only occurs when built-in instructions are carefully designed and sufficiently strong to initiate successful self-selection."
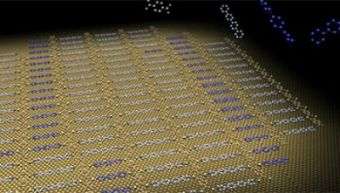
Scientists at the MPI directly observe the basic step of self-selection by imaging grid-like assemblies of molecules, which have sorted themselves by size. The features of the grid pattern are about one nanometer in size (0.000 000 001 meters), so small that they can only be imaged using state-of-the-art, ultra sensitive microscopy techniques. "Creating such miniscule architectures with features 50 000 times smaller than a hair is not a simple task," according to Dr. Steven Tait of the MPI. "Carving these nanometer structures with current technology would be inefficient and extremely expensive. Our strategy is to utilize instructed building blocks which can arrange themselves into desired structures."
The molecules are placed on ultra-clean metal surfaces and heated gently to enable motion, sorting, and organization. "The molecule movement on the copper surface is restricted to two-dimensions, but is still efficient enough to allow mixing of the molecules. By placing the molecules on a surface, we have the enormous advantage of being able to use specialized microscopes to ‚see’ the nanometer scale structures of the molecular assemblies," explains Alexander Langner, a graduate student at the MPI and first author of the study.
The study was conducted by Alexander Langner, Dr. Steven Tait, Dr. Nian Lin, and Prof. Dr. Klaus Kern of the Max Planck Institute for Solid State Research and Dr. Chandrasekar Rajadurai and Dr. Mario Ruben of the Karlsruhe Institute of Technology (KIT).
Professor Kern is the director of the Nanoscale Science Department at the MPI and leads a large research team conducting a wide range of studies related to the electronic, optical, and chemical properties of novel materials at the nanometer scale. Dr. Ruben is the leader of the research group "Functional Molecular Nanostructures" at the Institute of Nanotechnology in Karlsruhe and has a long-standing competence in the design and synthesis of instructed molecular components.
Source: Max-Planck-Gesellschaft