New software provides and overview of the big data of genome sequencing
- The amount of information that a genome researcher creates and which makes the basis of his scientific work has grown a million times during the last two decades. Today, the challenge does not consist in creating the data, but in exploring them and deducing meaningful conclusions. We believe that this analytical tool, which we have called "EaSeq" can help researchers in doing so, says Associate Professor Klaus Hansen
ChIP sequencing - an insight into the workflow of human cells
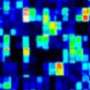
The EaSeq software has been developed for analysis of so called ChIP sequencing. DNA sequencing is used for mapping the sequence of the base pairs, which our DNA consists of, and ChIP sequencing is a derived method in which the sequences are used to determine the presence of different cell components in the genome at a given time.
Roughly speaking, ChIP sequencing can be compared to a microscope, which enables us to observe the presence of different cell components in the entire genome at a given time. The method is still quite young and holds the potential to be applied within many more scientific fields, which can benefit from understanding how healthy and pathological cells control and uses genes, says Associate Professor Mads Lerdrup
Better analytical tools means a broader range of applications
While ChIP sequencing has made it possible to produce enormous amounts of data very fast, the analysis of these data has - until now - been a tedious process. Most of the analytical software being used requires knowledge of computer programming and researchers have therefore been dependent on specialists in order to decode and analyze their data. EaSeq offers a far more visual and intuitive alternative, which makes it possible for biomedical researchers to study and test hypotheses using their own data. This means that instead of waiting for weeks for others to carry out an analysis, researchers will be able to perform the analyses themselves in a matter of hours.
Today, DNA sequencing is gaining ground within the clinical area where it is e.g. being used for diagnosis and targeting of treatment within the cancer area. The developers of EaSeq see similar perspectives for ChIP sequencing in the clinical work, and in that context strong analytical tools will be pivotal.
- The DNA sequence itself tells us very little about how cells actual decodes the DNA, and to understand this we need to map out which cell components are present in different parts of the genome at a specific time. It is our hope that we by increasing feasibility can enable researchers to faster uncover such knowledge and apply it clinically, says Associate professor Mads Lerdrup
Provided by University of Copenhagen