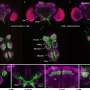
Study of complete RNA collection of fruit fly uncovers unprecedented complexity
(Phys.org) —Scientists from Indiana University are part of a consortium that has described the transcriptome of the fruit fly Drosophila melanogaster in unprecedented detail, identifying thousands of new genes, transcripts and proteins.
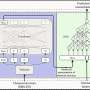
In the new work, published Sunday in the journal Nature, scientists studied the transcriptome—the complete collection of RNAs produced by a genome—at different stages of development, in diverse tissues, in cells growing in culture, and in flies stressed by environmental contaminants. To do so, they used contemporary sequencing technology to sequence all of the expressed RNAs in greater detail than ever before possible.
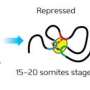
The paper shows that the Drosophila genome is far more complex than previously suspected and suggests that the same will be true of the genomes of other higher organisms. The paper also reports a number of novel, particular results: that a small set of genes used in the nervous system are responsible for a disproportionate level of complexity; that long regulatory and so-called "antisense" RNAs are especially prominent during gonadal development; that "splicing factors" (proteins that control the maturation of RNAs by splicing) are themselves spliced in complex ways; and that the Drosophila transcriptome undergoes large and interesting changes in response to environmental stresses.
The importance of Drosophila melanogaster as a model system cannot be overstated. Using it, the mechanisms of heredity were worked out about 100 years ago. Today, as biologists have developed increasing appreciation of how well genes and critical life processes are conserved over long evolutionary distances, flies have emerged as critical tools for understanding human biology and disease. Drosophila research is an area that has long had associations with IU, beginning with Nobel Laureate Herman J. Muller.

IU has 10 co-authors on the paper from the IU Bloomington College of Arts and Sciences' Department of Biology and the university's Center for Genomics and Bioinformatics. They are included among the 41 co-authors from 11 universities and institutes that are members of the National Human Genome Research Institute's Model Organism Encyclopedia of DNA Elements project, or modENCODE. Among the IU co-authors are Professor Emeritus of Biology Peter Cherbas, who helped manage the expansive project, and Distinguished Professor of Biology Thom Kaufman, who helped oversee design of the project and the production of biological samples.
"The modENCODE work is intended to provide a new baseline for research using Drosophila," Cherbas said. "The goal is to provide researchers working on particular processes with much of the detailed background information they would otherwise need to collect for themselves.
"As usual in science, we've answered a number of questions and raised even more. For example, we identified 1,468 new genes, of which 536 were found to reside in previously uncharacterized gene-free zones."
"We think these results could influence gene regulation research in all animals," Kaufman said. "This exhaustive study also identified a number of phenomena previously reported only in mammals, and that alone is really telling about the versatility of Drosophila melanogaster as a model organism. The new work provides a number of new potential uses for this powerful model system."
An example they pointed to was the perturbation experiments that identified new genes and transcripts. New genes were identified in experiments where adults were challenged with heat shock, cold shock, exposure to heavy metals, the drug caffeine and the herbicide paraquat, while larvae were treated with heavy metals, caffeine, ethanol or the insecticide rotenone.
Those environmental stresses resulted in small changes in expression level at thousands of genes; and in one treatment, four newly modeled genes were expressed altogether differently. In total, 5,249 transcript models for 811 genes were revealed only under perturbed conditions.
As did the flies in this new research, scientists who studied the Deepwater Horizon incident in the Gulf of Mexico found that marsh fishes responding to chronic hydrocarbon exposure had a number of expressional responses beyond the heat shock pathway, including the down regulation of genes encoding eggshell and yolk proteins as did the flies. To see this response overlap across phyla means the consortium may have identified a conserved metazoan stress response involving enhanced metabolism and the suppression of genes involved in reproduction.
More information: "Diversity and dynamics of the Drosophila transcriptome," James B. Brown, et al. Nature (2014) DOI: 10.1038/nature12962. Received 20 April 2013 Accepted 18 December 2013 Published online 16 March 2014
Journal information: Nature
Provided by Indiana University