October 16, 2012 report
Researchers discover a giant virus in an amoeba that contains a provirophage
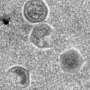
(Phys.org)—A team of researchers working out of the Centre National de la Recherche Scientifique in France has discovered a new giant virus living in an amoeba found in an eye patient's contact lens fluid. And as they describe in their paper published in the Proceedings of the National Academy of Sciences, they also found a new kind of virophage inside the virus which itself was harboring a previously unknown class of genetic parasite they've named transpovirons.
The researchers named the giant virus Lentille and found that it belongs to a group of giant viruses known as Mimiviridae. They named the virophage – a virus that infects other viruses – living inside the Lentille virus, Sputnik 2, (Russian for "fellow traveler"). Inside of the virophage they found fragments of DNA that didn't belong to the amoeba, the giant virus or the virophage, indicating that it was class of unknown genetic parasite that they've named transpovirons.
Sputnik 2 is just the fourth known virophage and is unique in that it can insert its DNA into its host genome, which might explain why the genomes of different giant viruses are so similar.
In looking closer at the virophage Sputnik 2, the team found fragments of DNA inside of it that didn't belong to any of the hosts; strands that outnumbered the giant viruses DNA by up to 14 times. Because there was no record of such fragments being observed before, the team gave them a name: transpovirons. As the name implies, the fragments were found able to jump in and out of host DNA in similar fashion to transposons.
The researchers believe transpovirons need giant viruses to replicate and are exceptionally good at reproducing and also appear to ride around in virophages. And because of the large variety of DNA found in them, the team believes they get their material from many different sources, similar in that respect to virophages, which other researchers have found contain the genomes of other giant viruses, bacteria or eukaryotic cells (simple microorganisms such as yeasts and fungi).
Because only a small amount of work has been focused on giant viruses, the team believes it's likely that other virophages and transposons will soon be discovered as more resources are dedicated to their study.
More information: Provirophages and transpovirons as the diverse mobilome of giant viruses, PNAS, Published online before print October 15, 2012, doi: 10.1073/pnas.1208835109
Abstract
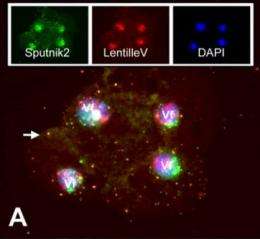
A distinct class of infectious agents, the virophages that infect giant viruses of the Mimiviridae family, has been recently described. Here we report the simultaneous discovery of a giant virus of Acanthamoeba polyphaga (Lentille virus) that contains an integrated genome of a virophage (Sputnik 2), and a member of a previously unknown class of mobile genetic elements, the transpovirons. The transpovirons are linear DNA elements of ∼7 kb that encompass six to eight protein-coding genes, two of which are homologous to virophage genes. Fluorescence in situ hybridization showed that the free form of the transpoviron replicates within the giant virus factory and accumulates in high copy numbers inside giant virus particles, Sputnik 2 particles, and amoeba cytoplasm. Analysis of deep-sequencing data showed that the virophage and the transpoviron can integrate in nearly any place in the chromosome of the giant virus host and that, although less frequently, the transpoviron can also be linked to the virophage chromosome. In addition, integrated fragments of transpoviron DNA were detected in several giant virus and Sputnik genomes. Analysis of 19 Mimivirus strains revealed three distinct transpovirons associated with three subgroups of Mimiviruses. The virophage, the transpoviron, and the previously identified self-splicing introns and inteins constitute the complex, interconnected mobilome of the giant viruses and are likely to substantially contribute to interviral gene transfer.
Journal information: Proceedings of the National Academy of Sciences
© 2012 Phys.org