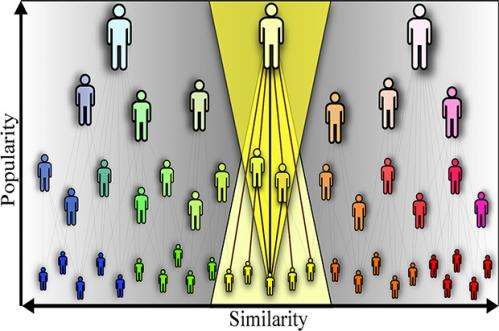
Popularity versus similarity: A balance that predicts network growth
(Phys.org)—Do you know who Michael Jackson or George Washington was? You most likely do: they are what we call "household names" because these individuals were so ubiquitous. But what about Giuseppe Tartini or John Bachar?
That's much less likely, unless you are a fan of Italian baroque music or free solo climbing.
In that case, you would have heard of Bachar just as likely as Washington. The latter was popular, while the former was not as popular but had interests similar to yours.
A new paper published this week in the science journal Nature by the Cooperative Association for Internet Data Analysis (CAIDA), based at the San Diego Supercomputer Center (SDSC) at the University of California, San Diego, explores the concept of popularity versus similarity, and if one more than the other fuels the growth of a variety of networks, whether it is the Internet, a social network of trust between people, or a biological network.
The researchers, in a study called Popularity Versus Similarity in Growing Networks, show for the first time how networks evolve optimizing a unique trade-off between popularity and similarity. They found that while popularity attracts new connections, similarity is just as attractive.
"Popular nodes in a network, or those that are more connected than others, tend to attract more new connections in growing networks," said Dmitri Krioukov, co-author of the Nature paper and a research scientist with SDSC's CAIDA group, which studies the practical and theoretical aspects of the Internet and other large networks. "But similarity between nodes is just as important because it is instrumental in determining precisely how these networks grow. Accounting for these similarities can help us better predict the creation of new links in evolving networks."
In the paper, Krioukov and his colleagues, which include network analysis experts from academic institutions in Cyprus and Spain, describe a new model that significantly increases the accuracy of network evolution prediction by considering the trade-offs between popularity and similarity. Their model describes large-scale evolution of three kinds of networks: technological (the Internet), social (a network of trust relationships between people), and biological (a metabolic network of the Escherichia coli, typically harmlessly found in the human gastrointestinal tract, though some strains can cause diarrheal diseases.)
The researchers write that the model's ability to predict links in networks may find applications ranging from predicting protein interactions or terrorist connections to improving recommender and collaborative filtering systems, such as Netflix or Amazon product recommendations.
"On a more general note, if we know the laws describing the dynamics of a complex system, then we not only can predict its behavior, but we may also find ways to better control it," added Krioukov.
In establishing connections in networks, nodes optimize certain trade-offs between the two dimensions of popularity and similarity, according to the researchers. "These two dimensions can be combined or mapped into a single space, and this mapping allows us to predict the probability of connections in networks with a remarkable accuracy," said Krioukov. "Not only can we capture all the structural properties of three very different networks, but also their large-scale growth dynamics. In short, these networks evolve almost exactly as our model predicts."
Many factors contribute to the probability of connections between nodes in real networks. In the Internet, for example, this probability depends on geographic, economic, political, technological, and many other factors, many of which are un-measurable or even unknown.
"The beauty of the new model is that it accounts for all of these factors, and projects them, properly weighted, into a single metric, while allowing us to predict the probability of new links with a high degree of precision," according to Krioukov.
More information: DOI: 10.1038/nature11459
Journal information: Nature
Provided by University of California - San Diego