Bacteria a potential threat to nuclear waste repositories

By interacting with the radioactive waste and the materials used to contain it, underground microorganisms may affect the safety of nuclear waste repositories, for better or for worse.
Underground, time appears to stand still. That is one of the reasons why deep geological formations are considered the safest place to dispose of nuclear waste. But now, scientists are finding out that human activities such as the excavation of tunnels can lead to a blooming of underground bacterial activity. In an ongoing research project, scientists from EPFL are cataloguing subterranean microbial life and studying its potential to affect the performance of the protective barriers - canisters, concrete and adjacent rock - that are used to contain nuclear waste.
Knowing what bacteria are present at such depths, what chemical transformations they are capable of, and even how they might evolve, have to be considered to assess the long-term safety of potential waste disposal sites. “We will use DNA sequencing and bio-informatics to identify the microbes trapped in the rock,” explains Rizlan Bernier-Latmani, head of EPFL’s Environmental Microbiology Laboratory and director of the project. The data will also provide insight on what types of molecules the microbes can produce using the substrates they grow on. “Given the long duration involved – several hundred thousand years, if the bacteria can do it, they will do it,” she says.
Bernier-Latmani has launched an experimental campaign hundreds of meters below Mont Terri, near St. Ursanne in the Swiss Jura. The Mont Terri site, which does not - and never will - hold nuclear waste, has become an international collaborative research platform looking at the suitability of argillaceous rock – in this case Opalinus Clay - for the disposal of radioactive waste. Opalinus Clay is one of the potential host rocks considered for nuclear waste disposal in Switzerland. Results from this experimental site will be largely transferrable to other similar sites.
Assessing the potential of microbes
Bernier-Latmani cites several bacterially-driven processes that have the potential to affect the safety of nuclear waste repositories. On the one hand, increased corrosion of metallic waste and containers and production of methane could both weaken the barriers that contain the radioactive waste. On the other hand, the microbes could consume gases that, over time, are likely to build up pressure in the sealed repository. Also, by altering the form of radioactive elements, their mobility could be reduced, efficiently immobilizing them in the bedrock.
“By furthering our understanding of what the microorganisms could do, we can take into account their activity in evaluating the long-term safety of the repository,” explains Bernier-Latmani. Using the latest DNA sequencing technology, she will be able to identify microbes even if they only represent a tenth of a percent of the microbial population. “It is very important to capture the low- abundance bacteria because they may include the genetic information that will allow them to thrive once the repository is sealed,” she says.
Watching the bacteria adapt in real-time
In a close-knit microbial community where different species coexist over millennia in a constantly changing environment, bacteria are likely to evolve and acquire new functions. In a process known as horizontal gene transfer, they can take up genes from other species. To get a clearer picture of how the bacteria might evolve over thousands of years, scientists have to identify the genes expressed by the microbial community as a whole.
How can we identify all of the bacterial species present in the clay deposits beneath Mont Terri? In the past, scientists did this by extracting bacteria from water samples and culturing them until they had a bacterial population large enough to analyze. But given that only about one percent of bacterial species can be cultured, their results hardly represented the true bacterial population.
Today, new techniques allow scientists to analyze bacterial populations without having to culture them in the lab. Using these techniques, the DNA of all microorganisms present in an environmental sample can be extracted, sequenced, and pieced together again using bioinformatics algorithms. This identifies all of the microbes and exhaustively catalogues their molecular machinery - the entire list of proteins that they can produce. Scientists can go even further and pinpoint the subset of these proteins that is actually in use at any given time, providing additional insight into the exact strategy the bacteria have adopted to survive, and how they might pose a threat.
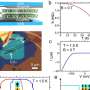
Bernier-Latmani and her team use a new kind of bioreactor, basically a bacterial aquarium that can be integrated right into the bedrock in tunnels hundreds of meters below ground. By providing a means of sampling the groundwater in the rock, these bioreactors let the researchers observe the bacteria’s growth and activity in their natural habitat.
For the time being, the EPFL researchers are interested in finding out how the bacteria respond to different diets they are likely to encounter in their underground habitat, and how their presence can affect the safety of repositories. The plan is to stimulate them by enriching the groundwater with different molecules, first with hydrogen to simulate the accumulation of the gas accompanying the corrosion of steel containers. All the while, they will have access to a number of parameters - pH, dissolved oxygen, and sulfate - in real-time. Using DNA sequencing techniques, they will be able to watch the bacteria adapt to their new environment.
It is still unknown whether the bugs are native or whether they colonized the bedrock during the excavation of the tunnels, but they are there, and they are active. Deep geological repositories have been selected as the most viable option for nuclear waste disposal, because, unlike on the surface of the earth, the bedrock appears to be frozen in time. But while geology may be close to standing still, human intrusion into the bedrock can bring underground biology back to its normal pace.
Provided by Ecole Polytechnique Federale de Lausanne