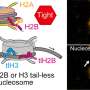
Nutrient supply after algal bloom determines the succession of the bacterial population

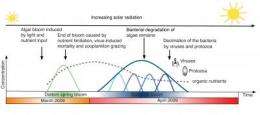
Algal blooms can considerably interfere with summer holidays by the sea. In the coastal zone of temperate regions a spring algal bloom is not a sign of excessive nutrient input, but most of all a consequence of the more intense solar irradiation in spring. When algal blooms end, the algae die and their remnants constitute an important nutrient supply for the whole ecosystem.
Researchers from the Max Planck Institute in Bremen, along with their co-authors from the University of Greifswald, the Jacobs University, and the Alfred Wegener Institute for Marine and Polar Research, examined an algal bloom in the North Sea and identified the microorganisms involved in the degradation of algal remnants. With their findings, the researchers discovered a solution for the so-called Plankton paradox: By specializing in different degradation processes, bacteria apparently occupy separate ecological niches in the sea.
For their analyses the scientists filtrated several hundreds of litres of seawater on a regular basis for almost a year off the station "Kabeltonne", a long-term station of the Biologische Anstalt Helgoland that is part of the Alfred Wegener Institute. "Pelagic microorganisms, the so called bacterioplankton, are critical for the breakdown of the dead algal biomass. Especially the dynamic succession in the bacterioplankton caught our attention. Specialized bacterial populations accompany different phases of the algal bloom", says Hanno Teeling from the Max Planck Institute As the scientists could show, processes within the bacterial population control the degradation of the algae.
His colleague Bernhard Fuchs who has been investigating the diversity and bacterioplankton composition for many years at the Max Planck Institute, adds: "For the first time we performed a high resolution analysis of the microbial community at genus level. We could not only identify the bacterial groups but also their functional tools, the enzymes that are involved in the breakdown of the algal bloom".
The results of the study may help the scientists to resolve the so-called plankton paradox: How can so many plankton species coexist in a seemingly homogeneous habitat without competing for nutrients in a way that eliminates certain species? Rudolf Amann, Director of the Max Planck Institute explains: "The secret at the level of the microorganisms is the heterogeneity of the microniches that the different groups inhabit. Thus, the specialized populations complement each other in the degradation of the organic matter."

The scientists used a novel combination of techniques for their analyses. They determined the identity of the microorganisms by CARD-FISH, an in situ technology that can be applied directly to environmental samples. Additionally, they probed the bacterial population during and after the algal bloom by short sequences of a phylogenetic marker gene (16S rRNA pyrotag analyses). "By using a combination of metagenome and metaproteome analyses we succeeded to detect the active key enzymes in complex environmental samples. This allows us to infer the role of the respective bacterial groups from their metabolic function", explains Thomas Schweder from the University of Greifswald. "This was only possibly by the computer-controlled integration of all data through bioinformatics,", states Frank Oliver Glöckner from the Max Planck Institute and Jacobs University.
In the early phase of the algal bloom the scientists encountered a variety of enzymes for the degradation of complex algal carbohydrates such as laminarin. At a later stage transport proteins for peptides, short protein units, and transporters for the growth limiting nutrient phosphate and simple sugar components dominated the enzymatic cocktail. Noteworthy was the high portion of certain transport proteins, the TonB-dependent transporters that can transport larger molecules directly into the interior of the cells.
This discovery may disprove the conventional acceptance that long-chained molecules need to be broken up into smaller components before the cell can take them up. The TonB-transporter may enable the Flavobacteria, one of the dominating bacterial groups, to couple the assimilation and degradation and thus to gain a competitive advance towards other bacterial groups. At the end of the bloom the bacteria increasingly produced sulfatases that cleave sulfate esters from algae carbohydrates hard to decompose and thus allow the complete degradation of these substances. Hence, the scientists discovered a bacterial population in the algae bloom that did not only differ in its composition but also in its function from the bacterial community found in crystal clear, remote open waters.
More information: Science 4 May 2012: vol. 336 no. 6081 pp. 608-611, DOI: 10.1126/science.1218344
Journal information: Science
Provided by Max Planck Society