1001 Genome-Project: On the way to a complete catalog of the Arabidopsis genome

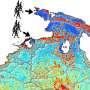
People can develop new technologies and animals may migrate to other regions. However, plants are tied to their location. Nevertheless, they have found ways to ensure their survival. This is the case for the plant Arabidopsis thaliana, which is found throughout the entire northern hemisphere. But how does this small, inconspicuous plant deal with all these different extremes?
In order to discover the whole-genome sequence variation, the 1001 Genomes Project was launched in 2008, with eleven research institutes participating worldwide. By investigating the genetic material of about one hundred strains of this plant from different geographical regions, researchers found a huge number of variations: in addition to millions of small differences that lead to a diversity of molecular gene products, they found hundreds of genes that are missing in some strains or have extra copies in others. It is probably this great flexibility within the genetic material that makes this plant particularly adaptable. In the medium term the complete catalog of the genome and gene product variation of a species can be applied to modern plant breeding.
Which genes and gene variants allow different individuals of one species to thrive under very different environmental conditions? The model plant for genetics, the thale cress, Arabidopsis thaliana, is especially well suited for the investigation of this question. It can deal with heat and drought in northern Africa as well as with cold in the central Asian highlands and temperate zones in Europe. Depending on the region it may display extensive foliage or appear small and fragile, yet it is always the same species. The answer lies without doubt in the diversity of its genetic material.
Detlef Weigel and Karsten Borgwardt from the Max Planck Institute for Developmental Biology, Gunnar Rätsch from the Friedrich Miescher Laboratory in Tübingen, and Karl Schmid of the University of Hohenheim have, together with an international team, sequenced and analyzed the genome of different Arabidopsis strains from all over Europe and Asia. To reveal the effect of geographic distance on the genes they selected plants from strains growing locally -- in the Swabian Neckar Valley -- as well as plants growing at opposite ends of the plant's distribution area, such as North Africa or Central Asia.
By sequencing nearly 100 genomes of different strains, the scientists hope to obtain a fundamental scientific understanding of evolution. The resulting information should pave the way for a new era of genetics in which alleles underpinning phenotypic diversity across the entire genome and the entire species can be identified. The scientists have found that thousands of proteins differ in their structure and function in the different Arabidopsis strains. In addition, they found several thousand cases of extra copies of genes, gene loss, as well as new genes that were previously only found in other plant species. "Our results show very impressively just how pronounced the genetic variability is," says Jun Cao from the Max Planck Institute for Developmental Biology and first author of one of the projects. Karl Schmid of the University of Hohenheim adds: "Adaptation through new mutations is very rare. More important is the recombination of already existing variants. With the information from more than a hundred genomes, not only can we make statements about these hundred individuals, but have thus laid the foundations to predict the genetic potential which could be realized by crossing particular individuals."
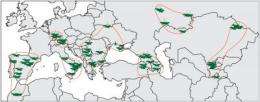
The geneticists working with Detlef Weigel, Karsten Borgwardt and Karl Schmid also found that the level of genetic variation differs widely between different regions. The researchers found the greatest genetic diversity in the Iberian Peninsula, where the plants have existed for a very long time. In Central Asia, which was only colonized after the last ice age, the Arabidopsis plants have relatively uniform genomes. Moreover, these populations have an above-average number of mutations that cause disadvantages for the plant, since protein functions are changed. Normally, natural selection removes these mutations over time, but in young emigrant populations they are enriched through cases of random evolution. "Figuring out how the plants and their genomes adapt to their environment is like a puzzle," says Jun Cao. "We need to collect all the pieces, before we can fit them together." The scientists have managed to create a nearly complete catalog of the genome variation of a species.
But how do these variations interact at the molecular level and what changes do they cause in the gene products? The computational biologist Gunnar Rätsch from the Friedrich Miescher Laboratory examined these questions in detail in a second study together with his international colleagues. They analyzed 19 strains of Arabidopsis with a particularly large genetic variability. These 19 individuals formed the basis of an artificial population of several hundred strains, created through multiple crosses such that different genome segments were shuffled systematically. The resulting individuals are ideally suited for examining gene interactions. The scientists studied the genome segments using novel analysis methods of analysis and discovered in detail how DNA is read in detail and how the intermediate stage of protein production, the RNA, is produced.
The researchers obtained detailed insight into the altered gene products arising from the various genomic variants. Depending on the genomic context some gene segments were either shut down or reactivated. "We can find a surprising number of changes affecting a single gene. However, they are often compensated for and therefore often have no significant effect on the gene products," says Gunnar Rätsch about the new results. The concepts, methods and platforms developed based on the genomic variation of Arabidopsis thaliana can also be used to study crop plants and for fast and accurate mapping of desirable characteristics. In addition, researchers can transfer this understanding about the influence of variation on gene products and their interactions to studies of the human genome.
These new projects should be viewed in the context of the 1001 Genomes Project, which was launched in 2008 at the Max Planck Institute for Developmental Biology and is being implemented through many individual projects in cooperation with ten other institutions worldwide. The aim is to analyze and compare the genes of 1001 different Arabidopsis strains. The goal of this large-scale project is to obtain fundamental insights into evolution, genetics and molecular mechanisms. Almost 500 different genomes have already been sequenced and analyzed at the different institutions. The data is being fed into a public database, which can be accessed not only by participants of the projects, but by all interested scientists.
More information:
Cao, J., Schneeberger, K., Ossowski, S., Günther, T., Bender, S., Fitz, J., Koenig, D., Lanz, C., Stegle, O., Lippert, C., Wang, X., Ott, F., Müller, J., Alonso-Blanco, C., Borgwardt, K., Schmid, K. J., and Weigel, D. (2011) Whole-genome sequencing of multiple Arabidopsis thaliana populations. Nature Genetics, doi: 10.1038/ng.911
Gan X., Stegle O., Behr J., Steffen J.G., Drewe P., Hildebrand K.L., Lyngsoe R., Schultheiss S.J., Osborne E.J., Sreedharan V.T., Kahles A., Bohnert R., Jean G., Derwent P., Kersey P., Belfield E.J., Harberd N.P., Kemen E., Toomajian C., Kover P.X., Clark R.M., Rätsch G., Mott R. (2011) Multiple reference genomes and transcriptomes for Arabidopsis thaliana. Nature, doi: 10.1038/nature10414
Schneeberger, K., Ossowski, S., Ott, F., Klein, J. D., Wang, X., Lanz, C., Smith, L. M., Cao, J., Fitz, J., Warthmann, N., Henz, S. R., Huson, D. H., and Weigel, D. (2011) Reference-guided assembly of four diverse Arabidopsis thaliana genomes. Proceedings of the National Academy of Sciences of USA 108, 10249-10254, doi: 10.1073/pnas.1107739108
Provided by Max-Planck-Gesellschaft