May 25, 2011 feature
Where no lab has gone before: Single-Molecule Electrokinetic Traps
(PhysOrg.com) -- To study the behavior of large protein complexes and long DNA chains in solution, researchers use so-called molecular traps. However, earlier traps have proven ineffective when working with small molecules due to the latter’s high diffusion. This limitation was first addressed through single-molecule immobilization techniques such as surface attachment and laser tweezers, but there were drawbacks: the former can disrupt biochemical structures, while the latter require molecules to be attached to large beads. A later trap developed at Stanford University used computer-based image capture and processing to track a single molecule’s Brownian motion, which it then cancels by applying variable voltage feedback. Now, however, Harvard University researchers have devised an Anti-Brownian ELectrokinetic (ABEL) trap that couples fluorescence microscopy to real-time electrokinetic feedback to trap any soluble fluorescence-capable molecule up to 800 times less massive than was previously possible.
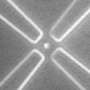
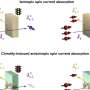
Developed at Harvard University by Prof. Adam E. Cohen in the Departments of Physics and of Chemistry and Chemical Biology, and Alex Fields, his student in the Biophysics Program, the ABEL trap works by following the Brownian motion of a particle, and then applying feedback forces to the particle to suppress this Brownian motion. The system uses fluorescence (from a dye molecule attached to that particle) to track the Brownian motion of the particle with a high degree of precision without damaging it.
Molecular traps face a basic challenge that has been historically difficult to overcome – namely, the differences in behavior between atoms in a low-temperature vacuum, which follow Newton’s laws of inertia and momentum, and those in solution. In the latter case, molecules collide every few picoseconds (as opposed to billions of times per second in a gas at atmospheric pressure, and only once every few seconds for typical atom trapping done at ultra-high vacuum), making it very difficult to track and analyze their positions and trajectories. This therefore requires a very different molecular trapping strategy.
While fluorescence has been employed in single-molecule imaging since the 1990s, these early systems required molecules to be immobilized on the surface of a slide using a chemical tether. Unfortunately, the tether often perturbed or modified the particle being trapped, so that there was no guarantee that it was behaving as it would if free in solution. Other early traps required molecules to be attached to small bead-like structures in order to be immobilized, which also often perturbed the molecule under study.
However, Cohen’s work with W. E. Moerner at Stanford University in 2005 led to traps that connected a CCD camera to a computer and determined the molecule's position via real-time image fitting. The computer then applied a time-varying feedback voltage to the solution so that the electrophoretic and electroosmotic drifts combined to cancel the Brownian motion.
Despite the significant progress made, the system’s speed was limited by software speed and the frame-rate of the camera. During his last year at Stanford, however, Cohen updated the trap with custom hardware that by having single-photon sensitivity allowed not only more precise measurement but the ability to determine where a photon had originated.
“The key to making ABEL work,” says Cohen, “is to do the feedback as quickly and accurately as possible. However, as we try to trap ever-smaller particles, this task becomes challenging for two reasons. First, smaller particles diffuse faster – the amount of diffusion is inversely proportional to the radius of the particle, so a 1 nm particle diffuses 10 times faster than a 10 nm particle. Second, smaller particles tend to be dimmer – and in the limit of having just one fluorescent dye molecule, we don't get very many photons from it. So in the end, we're trying to follow the motion of this incredibly quickly moving, dim object, and we need to do this with sub-millisecond resolution in time and micron-scale resolution in space. That's hard to do.”
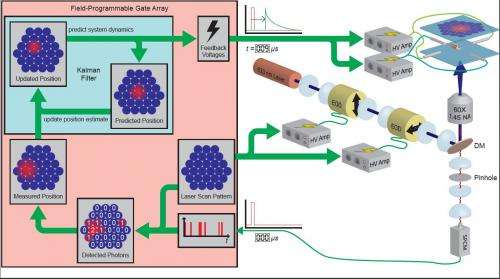
The primary innovation that allows the ABEL trap to trap single dye molecules is a statistically rigorous tracking algorithm that makes nearly optimal use of the information in every detected photon, which Fields designed and implemented in custom digital hardware (called a Field Programmable Gate Array, or FPGA). “The FPGA can run the algorithm tens of thousands of times per second,” explains Cohen, “so every time we detect a photon from a trapped molecule, the algorithm incorporates this information into its estimate of where the particle was, generates the appropriate feedback signals, and then waits until the next photon detection.”
The result is an imaging and detection system that performs the fastest and most sensitive tracking to date by combining all photon information in a statistically optimal way that generates the most likely estimate of the location of a photon’s source. Moreover, implementing the algorithm on the FPGA runs the algorithm in 9 μs, which is significantly less than the typical interval between photon emissions. (By way of comparison, the CCD camera-based system took 4.5 ms to process emitted photon data, and the algorithm prior to that developed by Fields required 25 μs.)
Cohen acknowledges that despite these advances, ABEL is not perfect. “One limitation to that ABEL can only trap for a few seconds because the oxygen-sensitive dye molecule is subject to laser-induced photobleaching. Optical excitation has a small probability of resulting in a photochemical change that disrupts the dye molecule.” However, he adds that photobleaching can be minimized by adding chemicals that consume oxygen, as well as other chemicals that decrease its impact, such as antioxidants and free- radical scavengers.
Going forward, Cohen is interested in layering other kinds of spectroscopy on top of the trapping. “We want to shine different colors of light onto the trapped molecule, and so get more information out of the photons – their polarization, wavelength, and precise timing – that the molecule emits. This additional information will give us a more detailed picture of what the one molecule is doing inside the trap”
Cohen also is investigating the addition of various fluidics to facilitate the inflow and outflow of different reagents. “It would be great if we could trap an enzyme, say, and then flow in substrate or ATP, and see how the dynamics of the enzyme change.”
Regarding applications, Cohen is hoping in the near term to study the dynamics of short pieces of DNA and DNA-protein interactions. “DNA has been incredibly well studied, of course; but there are still really fundamental and important things we don't know,” he points out. “For example, we don't know what happens to DNA if you bend it very sharply, or how the mechanical properties of DNA depend on its underlying sequence. These questions are important to the function of DNA in a cell, because DNA in a cell is often highly bent around histones or by DNA-binding proteins. We also don't fully understand how DNA-binding proteins find their specific binding sites on the molecule. It's possible that the proteins are probing the local mechanical properties of the molecule as part of their search.”
In the longer term, the team would like to study the internal dynamics of a wide range of proteins and molecular machines. “Now that we can trap nearly any molecule without tethering it to a surface, we can hope to look at the dynamics of many individual molecules that thus far have been impossible to study at the single-molecule level.”
More information:
• Electrokinetic trapping at the one nanometer limit, PNAS Published online before print May 11, 2011, DOI:10.1073/pnas.1103554108
• Cohen Lab at Harvard University
Copyright 2011 PhysOrg.com.
All rights reserved. This material may not be published, broadcast, rewritten or redistributed in whole or part without the express written permission of PhysOrg.com.