This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
The 'jigglings and wigglings of atoms' reveal key aspects of COVID-19 virulence evolution
Richard Feynman famously stated, "Everything that living things do can be understood in terms of the jigglings and wigglings of atoms." This week, Nature Nanotechnology features a study that sheds new light on the evolution of the coronavirus and its variants of concern by analyzing the behavior of atoms in the proteins at the interface between the virus and humans.
The paper, titled "Single-molecule force stability of the SARS-CoV-2–ACE2 interface in variants-of-concern," is the result of an international collaborative effort among researchers from six universities across three countries.
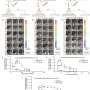
The study introduces significant insights into the mechanical stability of the coronavirus, a key factor in its evolution into a global pandemic. The research team employed advanced computational simulations and magnetic tweezers technology to explore the biomechanical properties of biochemical bonds in the virus. Their findings reveal critical distinctions in the mechanical stability of various virus strains, highlighting how these differences contribute to the virus's aggressiveness and spread.
As the World Health Organization reports nearly 7 million deaths worldwide from COVID-19, with more than 1 million in the United States alone, understanding these mechanics becomes crucial for developing effective interventions and treatments. The group emphasizes that comprehending the molecular intricacies of this pandemic is key to shaping our response to future viral outbreaks.
Delving deeper into the study, the Auburn University team, led by Prof. Rafael C. Bernardi, Assistant Professor of Biophysics, along with Dr. Marcelo Melo and Dr. Priscila Gomes, played a pivotal role in the research by leveraging powerful computational analysis. Utilizing NVIDIA HGX-A100 nodes for GPU computing, their work was essential in unraveling complex aspects of the virus's behavior.
Prof. Bernardi, an NSF Career Award recipient, collaborated closely with Prof. Gaub from LMU, Germany, and Prof. Lipfert from Utrecht University, The Netherlands. Their collective expertise spanned various fields, culminating in a comprehensive understanding of the SARS-CoV-2 virulence factor. Their research demonstrates that the equilibrium binding affinity and mechanical stability of the virus–human interface are not always correlated, a finding crucial for comprehending the dynamics of viral spread and evolution.
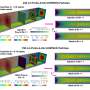
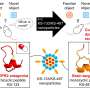
Additionally, the team's use of magnetic tweezers to study the force-stability and bond kinetics of the SARS-CoV-2:ACE2 interface in various virus strains provides new perspectives on predicting mutations and adjusting therapeutic strategies. The methodology is unique because it measures how strongly the virus binds to the ACE2 receptor, a key entry point into human cells, under conditions that mimic the human respiratory tract.
The group found that while all the major COVID-19 variants (like alpha, beta, gamma, delta, and omicron) bind more strongly to human cells than the original virus, the alpha variant is particularly stable in its binding. This might explain why it spread so quickly in populations without prior immunity to COVID-19. The results also suggest that other variants, like beta and gamma, evolved in a way that helps them evade some immune responses, which might give them an advantage in areas where people have some immunity, either from previous infections or vaccinations.
Interestingly, the delta and omicron variants, which became dominant worldwide, show traits that help them escape immune defenses and possibly spread more easily. However, they don't necessarily bind more strongly than other variants. Prof. Bernardi says, "This research is important because it helps us understand why some COVID-19 variants spread more quickly than others. By studying the virus's binding mechanism, we can predict which variants might become more prevalent and prepare better responses to them."
This research emphasizes the importance of biomechanics in understanding viral pathogenesis and opens new avenues for scientific investigation into viral evolution and therapeutic development. It stands as a testament to the collaborative nature of scientific research in addressing significant health challenges.
More information: Magnus S. Bauer et al, Single-molecule force stability of the SARS-CoV-2–ACE2 interface in variants-of-concern, Nature Nanotechnology (2023). DOI: 10.1038/s41565-023-01536-7. www.nature.com/articles/s41565-023-01536-7
Journal information: Nature Nanotechnology
Provided by Auburn University